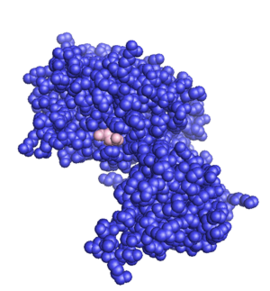
Function
PTEN protein acts as a phosphatase to dephosphorylate phosphatidylinositol (3,4,5)-trisphosphate (PtdIns (3,4,5)P3 or PIP3). PTEN specifically catalyses the dephosphorylation of the 3` phosphate of the inositol ring in PIP3, resulting in the biphosphate product PIP2 (PtdIns(4,5)P2). This dephosphorylation is important because it results in inhibition of the Akt signaling pathway, which plays an important role in regulating cellular behaviors such as cell growth, survival, and migration.
PTEN also has weak protein phosphatase activity, but this activity is also crucial for its role as a tumor suppressor. PTEN's protein phosphatase activity may be involved in the regulation of the cell cycle, preventing cells from growing and dividing too rapidly.[8] There have been numerous reported protein substrates for PTEN, including IRS1[10] and Dishevelled.[11]
PTEN is one of the targets for drug candidates such as the oncomiR, MIRN21.
Structure
The structure of the core of PTEN (solved by X-ray crystallography, see figure to the upper right[5]) reveals that it consists primarily of a phosphatase domain, and a C2 domain: the phosphatase domain contains the active site, which carries out the enzymatic function of the protein, while the C2 domain binds the phospholipid membrane. Thus PTEN binds the membrane through both its phosphatase and C2 domains, bringing the active site to the membrane-bound PIP3 to dephosphorylate it.
The two domains of PTEN, a protein tyrosine phosphatase domain and a C2 domain, are inherited together as a single unit and thus constitute a superdomain, not only in PTEN but also in various other proteins in fungi, plants and animals, for example, tensin proteins and auxilin.[12]
The active site of PTEN consists of three loops, the TI Loop, the P Loop, and the WPD Loop, all named following the PTPB1 nomenclature.[5] Together they form an unusually deep and wide pocket which allows PTEN to accommodate the bulky phosphatidylinositol 3,4,5-trisphosphate substrate. The dephosphorylation reaction mechanism of PTEN is thought to proceed through a phosphoenzyme intermediate, with the formation of a phosphodiester bond on the active site cysteine, C124.
Not present in the crystal structure of PTEN is a short 10-amino-acid unstructured region N-terminal of the phosphatase domain (from residues 6 to 15), known variously as the PIP2 Binding Domain (PBD) or PIP2 Binding Motif (PBM)[13][14][15] This region increases PTEN's affinity for the plasma membrane by binding to Phosphatidylinositol 4,5-bisphosphate, or possibly any anionic lipid.
Also not present in the crystal structure is the intrinsically disordered C-terminal region (CTR) (spanning residues 353–403). The CTR is constitutively phosphorylated at various positions that effect various aspects of PTEN, including its ability to bind to lipid membranes, and also act as either a protein or lipid phosphatase.[16][17]
Additionally, PTEN can also be expressed as PTEN-L[18] (known as PTEN-Long, or PTEN-α[19]), a leucine initiator alternative start site variant, which adds an additional 173 amino acids to the N-terminus of PTEN. The exact role of this 173-amino acid extension is not yet known, either causing PTEN to be secreted from the cell, or to interact with the mitochondria. The N-terminal extension has been predicted to be largely disordered,[20] although there is evidence that there is some structure in the last twenty amino acids of the extension (most proximal to the start methionine of PTEN).[17]
Clinical significance
Cancer
PTEN is one of the most commonly lost tumor suppressors in human cancer; in fact, up to 70% of men with prostate cancer are estimated to have lost a copy of the PTEN gene at the time of diagnosis.[21] A number of studies have found increased frequency of PTEN loss in tumours which are more highly visible on diagnostic scans such as mpMRI, potentially reflecting increased proliferation and cell density in these tumours.[22]
During tumor development, mutations and deletions of PTEN occur that inactivate its enzymatic activity leading to increased cell proliferation and reduced cell death. Frequent genetic inactivation of PTEN occurs in glioblastoma, endometrial cancer, and prostate cancer; and reduced expression is found in many other tumor types such as lung and breast cancer. Furthermore, PTEN mutation also causes a variety of inherited predispositions to cancer.
Non-cancerous neoplasia
Researchers have identified more than 70 mutations in the PTEN gene in people with Cowden syndrome.[citation needed] These mutations can be changes in a small number of base pairs or, in some cases, deletions of a large number of base pairs.[citation needed] Most of these mutations cause the PTEN gene to make a protein that does not function properly or does not work at all. The defective protein is unable to stop cell division or signal abnormal cells to die, which can lead to tumor growth, particularly in the breast, thyroid, or uterus.[23]
Mutations in the PTEN gene cause several other disorders that, like Cowden syndrome, are characterized by the development of non-cancerous tumors called hamartomas. These disorders include Bannayan–Riley–Ruvalcaba syndrome and Proteus-like syndrome. Together, the disorders caused by PTEN mutations are called PTEN hamartoma tumor syndromes, or PHTS. Mutations responsible for these syndromes cause the resulting protein to be non-functional or absent. The defective protein allows the cell to divide in an uncontrolled way and prevents damaged cells from dying, which can lead to the growth of tumors.[23]
Brain function and autism
Defects of the PTEN gene have been cited to be a potential cause of autism spectrum disorders.[24]
When defective, PTEN protein interacts with the protein of a second gene known as Tp53 to dampen energy production in neurons. This severe stress leads to a spike in harmful mitochondrial DNA changes and abnormal levels of energy production in the cerebellum and hippocampus, brain regions critical for social behavior and cognition. When PTEN protein is insufficient, its interaction with p53 triggers deficiencies and defects in other proteins that also have been found in patients with learning disabilities including autism.[24] People with autism and PTEN mutations may have macrocephaly (unusually large heads).[25]
Patients with defective PTEN can develop cerebellar mass lesions called dysplastic gangliocytomas or Lhermitte–Duclos disease.[23]
Cell regeneration
PTEN's strong link to cell growth inhibition is being studied as a possible therapeutic target in tissues that do not traditionally regenerate in mature animals, such as central neurons. PTEN deletion mutants have recently[26] been shown to allow nerve regeneration in mice.[27][28]
As a drug target
PTEN inhibitors
Bisperoxovanadium compounds may have a neuroprotective effect after CNS injury.[29] PTEN is inhibited by sarcopoterium.[30]
Cell lines
Cell lines with known PTEN mutations include: