| This is an archive of past discussions. Do not edit the contents of this page. If you wish to start a new discussion or revive an old one, please do so on the current talk page. |
| Archive 1 | Archive 2 | Archive 3 | Archive 4 |
Rat genome database listed at Requested moves

A requested move discussion has been initiated for Rat genome database to be moved to Rat Genome Database. This page is of interest to this WikiProject and interested members may want to participate in the discussion here. —RMCD bot 16:31, 26 July 2015 (UTC)
- To opt out of RM notifications on this page, transclude ((bots|deny=RMCD bot)), or set up Article alerts for this WikiProject.
This abandoned draft needs to be merged into the existing 45,X/46,XY mosaicism stub, before the draft is deleted as stale. The draft contains far more detail than the mainspace stub. Roger (Dodger67) (talk) 14:43, 24 August 2015 (UTC)
- Anyone? Roger (Dodger67) (talk) 14:18, 25 August 2015 (UTC)
- I have a slow morning. I'll take it. Cheers, BatteryIncluded (talk) 15:16, 25 August 2015 (UTC)
- Note: A synonym of the disease is Mixed gonadal dysgenesis, so we'll need to merge this article too. BatteryIncluded (talk) 16:09, 25 August 2015 (UTC)
- I have a slow morning. I'll take it. Cheers, BatteryIncluded (talk) 15:16, 25 August 2015 (UTC)
- @Dodger67: I merged the Draft:45,X/46,XY mosaicism and Mixed gonadal dysgenesis into 45,X/46,XY mosaicism. I also did a small expansion. I think both the draft and 'Mixed gonadal dysgenesis' are ready to be redirected and deleted. Cheers, BatteryIncluded (talk) 17:17, 25 August 2015 (UTC)
- @BatteryIncluded: Thanks, the finishing touches have been done; redirects, class parameters, removing merge templates. Roger (Dodger67) (talk) 17:51, 25 August 2015 (UTC)
- @Dodger67: I merged the Draft:45,X/46,XY mosaicism and Mixed gonadal dysgenesis into 45,X/46,XY mosaicism. I also did a small expansion. I think both the draft and 'Mixed gonadal dysgenesis' are ready to be redirected and deleted. Cheers, BatteryIncluded (talk) 17:17, 25 August 2015 (UTC)
Hello, genetics experts. Here's an old draft that will soon be deleted as stale unless someone takes an interest in it. Is this a notable topic?—Anne Delong (talk) 15:01, 24 August 2015 (UTC)
- I have merged the usable content into the stop codon article, as suggested by the AFC reviewer. I think the draft can now be deleted. Looie496 (talk) 15:32, 24 August 2015 (UTC)
- Thanks, Looie496. Rather than delete the draft, I've redirected it and moved it to Nonstop mutation. That way, if it is ever desirable for the topic to have its own article, the text in the history can be restored and improved without the need for a split.—Anne Delong (talk) 22:09, 24 August 2015 (UTC)
GMO Wheat
The GMO Wheat article is out of date. it looks like all references are from 2013.96.40.168.151 (talk) 20:40, 23 June 2015 (UTC)
Using raw data from peer reviewed papers
I've started a discussion at Wikipedia talk:Identifying reliable sources#Using data from academic papers on genetics. Doug Weller (talk) 15:43, 5 August 2015 (UTC)
Exon article
Hi there, I'm new to this project and to editing wikipedia articles in general. When I read through the exon article today, I noticed that it's rather short and that the introductory sentence is quite misleading, as was already discussed in the Talk page. However, I don't have a final good idea how to change the introductory sentence to make it more precise. In my opinion, exons encode genes (or rather, proteins/mRNA), and they are also encoded by the gene itself, since they are a part of it... So I thought, maybe someone else could give their opinion, as to how this should be settled? I think it's rather sad that this important article is having such issues and that it hasn't grown yet. — Preceding unsigned comment added by Ilikelifesciences (talk • contribs) 09:12, 2 September 2015 (UTC)
- @Ilikelifesciences: Hello, and welcome to WP:MCB! I notice the Exon article doesn't have any references in the lead. I reckon that the firswt step would have to be to read a couple of references to see how they define it. There are a couple of reviews cited lower down the page, or I'm sure google scholar can suggest something to read. It must be a very well reviewed topic by now. Another source of info could be the Molecular Biology of the Cell online textbook. I reckon that the page probably needs much improvement of the Function section and new sections on Origins & evolution, Occurence, and Annotation. I'm working on other pages at the moment, but would be happy to chip in if you want to work on it! T.Shafee(Evo﹠Evo)talk 11:10, 2 September 2015 (UTC)
- What was written in the lead was so garbled that it was useless. I have, I believe, improved it. Maproom (talk) 22:44, 2 September 2015 (UTC)
Category:Sequenced genomes
Category:Sequenced genomes, which is within the scope of this WikiProject, has been nominated for renaming to "Category:Lists of sequenced genomes" with the deletion of all sub-categories. If you would like to participate in the discussion, you are invited to add your comments at the category's entry on the Categories for discussion page. Thank you. RevelationDirect (talk) 02:18, 10 October 2015 (UTC)
Review of new edit to Indo-Aryan peoples requested
It would be helpful to have other views on the genetics related edit at [1] - see also the talk page for some context. Thanks. Doug Weller (talk) 19:21, 11 October 2015 (UTC)
- Genetics and archaeogenetics of South Asia is also pretty dense for the average reader.Doug Weller (talk) 05:02, 12 October 2015 (UTC)
Not sure what to make of this article. The lead says "Genetic anthropology (Human genetics) is a branch of human genetics" and then says it's a "field of anthropology." I removed an unsourced section which said:
"There seems to be a divide in the academic community as to what is Genetic Anthropology and what is Molecular Anthropology. There are many universities that offer degrees in Genetic Anthropology or Anthropological Genetics, while others offer degrees in Molecular Anthropology, but all are considered to be under the umbrella of Evolutionary Anthropology. In most universities genetic anthropology is taught in conjunction with other anthropological areas such as paleoanthropology, primatology, human biology, Bioarchaeology and osteology."
IMHO this needs to be redirect or rewritten. Crossposting to Wikipedia talk:WikiProject Anthropology Doug Weller (talk) 10:48, 22 October 2015 (UTC)
- Ok, you can call me a reductionist bigot, but what the article says to me is "Genetic anthropology is a wishy-washy attempt by humanities students to study human genetics". Maproom (talk) 13:55, 22 October 2015 (UTC)
- Yes, it looks like a simplified course on population genetics for anthropologists. BatteryIncluded (talk) 14:14, 22 October 2015 (UTC)
- If that is right, and it is the name of a course or courses, rather than a field of scientific investigation, it probably shouldn't be the subject of an article. Maproom (talk) 16:59, 22 October 2015 (UTC)
- Yes, it looks like a simplified course on population genetics for anthropologists. BatteryIncluded (talk) 14:14, 22 October 2015 (UTC)
DNA Extraction
Hello I'm new to wikipedia and this project and would like to suggest a revision on the DNA Extraction article. My co-wokers and I are experts in salvia DNA extraction and would like to add more specifics about types of extraction into the current article. I suggest changing the "Special Types of DNA Extraction" to "Methods of DNA Extraction" or "Types of DNA Extraction" I would like to collaborate with others on the project who could give specifics about other methods of DNA extraction such as blood, microorganisms, ancient... etc. We'd be happy to write out the specifics of salvia DNA extraction and create a template for others to use. Let me know what you think of this idea and if you are someone who may be able to help. Thank you! Amh97 (talk) 15:35, 30 October 2015 (UTC) 'd
- Hi Amh97, your desire to help is much appreciated. I invite you to contribute your expertise/knowledge to the article in anyway you see fit just as long as the information is verifiable and cites reliable sources. Bear in mind that any methodology that your lab may have adopted and refined may constitute original research. So just be careful. Happy editing. Wisdom89 (T / C) 15:54, 30 October 2015 (UTC)
- One other thing that I should have made clear, you are under no obligation to ask permission to edit. This is an open encyclopedia : ) Wisdom89 (T / C) 15:56, 30 October 2015 (UTC)
- Amh97 Hello and Welcome to Wikipedia! As Wisdom89 noted, you can jump in any time to edit. The learning curve using Wikipedia is not steep, just make sure you cite reliable references by inserting the bibliography or web site in between this code at the end of important sentences: <ref> http: xyz </ref> Alternatively, you can use this machine to help you format the cited reference: https://tools.wmflabs.org/makeref/
- I will keep an eye on the DNA extraction article to assist you with formatting and miscellaneous. Also, keep in mind that the subject is in general DNA extraction procedures, so do not go heavily on methods for salvia. Cheers, BatteryIncluded (talk) 16:28, 30 October 2015 (UTC)
Draft:Hox Genes in Amphibians and Reptiles
Dear genetics experts: This old draft has references, but they are not on line. The text read like a journal article. I haven't been able to find any copyright violation, but I don't have access to the right journals. Is this page worth keeping, or should it be deleted as a stale draft (which it can be if no one edits it).—Anne Delong (talk) 00:35, 3 November 2015 (UTC)
- I'll give it a shot. Cheers, BatteryIncluded (talk) 01:04, 3 November 2015 (UTC)
- Thanks, BatteryIncluded. I have postponed its deletion to give you time to fix it up.—Anne Delong (talk) 07:53, 3 November 2015 (UTC)
- Overhauled to Wikipedia standards - hopefully. Posted now live at Hox genes in amphibians and reptiles. Please review at will. Cheers, BatteryIncluded (talk) 18:50, 3 November 2015 (UTC)
- I will make sure the next draft lands on your desk. Thank you for your help. Cheers, BatteryIncluded (talk) 02:07, 4 November 2015 (UTC)
- BatteryIncluded, thanks for improving that page; just one point for next time: please move the page to mainspace instead of copying it; I had to do a historymerge. However, at least I knew how to do that, whereas I know little about genetics, so your help is greatly appreciated.—Anne Delong (talk) 03:08, 4 November 2015 (UTC)
- I will make sure the next draft lands on your desk. Thank you for your help. Cheers, BatteryIncluded (talk) 02:07, 4 November 2015 (UTC)
- Overhauled to Wikipedia standards - hopefully. Posted now live at Hox genes in amphibians and reptiles. Please review at will. Cheers, BatteryIncluded (talk) 18:50, 3 November 2015 (UTC)
- Thanks, BatteryIncluded. I have postponed its deletion to give you time to fix it up.—Anne Delong (talk) 07:53, 3 November 2015 (UTC)
DNA listed at Requested moves

A requested move discussion has been initiated for DNA to be moved to Deoxyribonucleic acid. This page is of interest to this WikiProject and interested members may want to participate in the discussion here. —RMCD bot 20:29, 16 November 2015 (UTC)
- To opt out of RM notifications on this page, transclude ((bots|deny=RMCD bot)), or set up Article alerts for this WikiProject.
Articles by @Rcrzarg:
Can someone with non-coding RNA knowledge look at the article creations of Rcrzarg to see if they are salvageable. This is not my area, but to me they look like original research. For example, from Αr35 RNA: "To identify binding sites for other known transcription factors we used the fasta sequences provided by RegPredict" "This [Covariance Model] was used in a further search for new members of the αr35 family in the existing bacterial genomic databases". Thanks, 109.79.192.231 (talk) 12:49, 13 January 2016 (UTC)
Discussion about generally considering articles from predatory publishers unreliable
There is a discussion here if that topic is of interest. It has been going on since Feb 26, but just wanted to make sure folks here are aware of it. Jytdog (talk) 18:06, 4 March 2016 (UTC)
Suggestion: Create Infobox for "Repeated Sequences"
I'm trying to improve some of the articles on repeated sequences and thought it might be helpful if there was an infobox that could be used. There is already the very nice Template for Repeated sequences, but maybe some kind of Infobox would be nice? However, this should of course not overlap with the already existing template. Maybe it could contain information on the repeated sequence's propagation mechanism (copy/cut paste) and it's prevalence in specific organisms? Looking forward to any suggestions! Ilikelifesciences (talk) 11:50, 5 April 2016 (UTC)
Palomino
Is the color, as described in the lead of Palomino horse, created the same way in the Palomino rabbit? Many thanks. Anna Frodesiak (talk) 22:30, 7 April 2016 (UTC)
- The Palomino horse is heterozygous for the one relevant gene. As far as I can tell from the Palomino rabbit article, the rabbit isn't (necessarily) heterozygous, it's just a similar colour to the horse. Maproom (talk) 23:20, 7 April 2016 (UTC)
- Thank you kindly, Maproom. :) Anna Frodesiak (talk) 23:21, 7 April 2016 (UTC)
- I added a bit of a disclaimer based on what you said. Do you think it's okay? Anna Frodesiak (talk) 23:29, 7 April 2016 (UTC)
- I think it's okay in that I don't disagree with it. But it's a bit long, and the final phrase is just saying "we don't know whether this is the case". Maybe instead "As with the palomino horse, the rabbit gets its name from its color,[1][2] though the genes involved are probably different." Maproom (talk) 19:19, 9 April 2016 (UTC)
- Well put, Maproom. Article updated. Thank you so much! :) Anna Frodesiak (talk) 22:29, 9 April 2016 (UTC)
- I think it's okay in that I don't disagree with it. But it's a bit long, and the final phrase is just saying "we don't know whether this is the case". Maybe instead "As with the palomino horse, the rabbit gets its name from its color,[1][2] though the genes involved are probably different." Maproom (talk) 19:19, 9 April 2016 (UTC)
Please see Talk:Tinman gene about the wholesale replacement of the existing article content with the content from a draft. I object to such overwriting of good content and prefer to se a proper merge 9f the best of the old and new content. The editor who did the replacement is new so needs assistance from experienced editors familiar with the subject. Roger (Dodger67) (talk) 08:44, 14 April 2016 (UTC)
- P.S. The draft is at Draft:Tinman (Nkx2-5) gene -- Roger (Dodger67) (talk) 09:33, 14 April 2016 (UTC)
Adding to the list of bacterial genomes
I noticed that Sorangium cellulosum was missing, so I added it to List of sequenced bacterial genomes#Delta/epsilon subdivisions. It's interesting as the largest known bacterial genome (at least up to the date of publication, and none on the list are larger). Unfortunately I'm not knowledgeable about genetics or microbiology -- would someone check this over and make sure it looks right?
In particular, do I have the right strain and gene count? Is my citation OK?
CRGreathouse (t | c) 16:30, 19 April 2016 (UTC)
See discussion importing content from protein database into WP
Here: Wikipedia_talk:WikiProject_Molecular_and_Cell_Biology#Transporter_classification_database Jytdog (talk) 22:12, 24 April 2016 (UTC)
Gene drives
Due to its major implications and because it's actively developing, gene drives should be presented in more detailed. Also the page should be rated "start" and "top" (or at least "high") importance in my opinion. (Tjulou (talk) 05:36, 15 May 2016 (UTC))
- @Tjulou: I've changed the class to "start" but the importance to "mid" for the moment (I don't know a huge amount about the topic) since I think it's not hugely common. I'll let someone with more knowledge bump it up to "high" if appropriate. T.Shafee(Evo﹠Evo)talk 07:10, 15 May 2016 (UTC)
Auto-assessment of article classes
Following a recent discussion at WP:VPR, there is consensus for an opt-in bot task that automatically assesses the class of articles based on classes listed for other project templates on the same page. In other words, if WikiProject A has evaluated an article to be C-class and WikiProject B hasn't evaluated the article at all, such a bot task would automatically evaluate the article as C-class for WikiProject B.
If you think auto-assessment might benefit this project, consider discussing it with other members here. For more information or to request an auto-assessment run, please visit User:BU RoBOT/autoassess. This is a one-time message to alert projects with over 1,000 unassessed articles to this possibility. ~ RobTalk 22:28, 3 June 2016 (UTC)
Merge discussion
A merge discussion is taking place here to determine whether Gene and Genomic organization should be merged. Seems important. Best Regards,
- Barbara (WVS) (talk) 10:58, 30 June 2016 (UTC)
A user (Yahadzija) has recently made a flurry of changes to the Epistasis page, splitting most of it off into Non-allelic gene interaction and adding their own content to expand the old epistasis page. I disagree with the distinction that they've tried to draw, however other genetics editors' opinions would be very helpful to see if I'm missing something. T.Shafee(Evo﹠Evo)talk 00:30, 3 July 2016 (UTC)
- I'm not too familiar with the jargon of the field, but I haven't heard the term "non-allelic gene interaction" thrown around (and a quick pubmed search for the term didn't turn up much). It's not clear from the articles as they stand now how "non-allelic gene interaction" differs from epistasis (or rather how non-allelic gene interaction is a category that epistasis falls into), but I'm happy to be educated by any more familiar with the topic. Ajpolino (talk) 05:11, 3 July 2016 (UTC)
- The Epistasis article treats "epistasis" as a special case of non-allelic gene interaction, with an allele at one locus suppressing the effect of the alleles present at another. The Non-allelic gene interaction article makes extensive use of the word "epistasis" in senses incompatible with that definition. This is a mess, which could be cleared up by merging the two articles. Maproom (talk) 08:04, 3 July 2016 (UTC)
WikiFactMine: Proposed Wiki Grant for genetic information in Wikidata
We are proposing a Wikimedia Project Grant (WikiFactMine) which will automatically scan the daily peer-reviewed scientific literature (up to 10000 articles /day) and extract genes (definable in Wikidata). We concentrate on genes which have high precision and recall, but with Wikidata-based dictionaries this can be extended to other well defined entities (e.g. cell types or diseases). These, with their citations, are then offered to Wikidata editors for potential inclusion/update. These can also alert Wikipedia editors to new citations. The project includes a Wikimedian in residence in the University of Cambridge. We'd be grateful for comments, endorsement and offers of help.Petermr (talk) 10:37, 2 August 2016 (UTC)
This project's feedback would be appreciated in this discussion, as this could greatly (and positively) affect biological citations! Headbomb {talk / contribs / physics / books} 21:53, 7 September 2016 (UTC)
This new article needs some help, the principal author, Dpaulbick, is new to WP. Roger (Dodger67) (talk) 17:45, 12 January 2017 (UTC)
- That article reads like a rambling incoherent essay, sometimes with little relationship to its lead. Its point seems to be that there are genes that we don't know what they do. Maproom (talk) 18:13, 12 January 2017 (UTC)
- The subject of this article I think is more subtle. We know what many of these genes do, but we don't know the impact of mutations in these genes have on patients. This is more of a medical than scientific article. This should be explained better in the lead. Without further clarification, the article is very confusing. Boghog (talk) 19:54, 12 January 2017 (UTC)
- The main source in this article is PMID 25741868. In closer reading of the source, it appears what is really meant by gene of uncertain significance (GUS) is gene variant of uncertain significance (GVUS). Boghog (talk) 20:02, 12 January 2017 (UTC)
- Aren't gene variants normally known as "alleles"? Maproom (talk) 22:03, 12 January 2017 (UTC)
- Yes, but variant indicates an allele different from the reference genome, as in a VCF file. In clinical sequencing, we talk of variants and variants of unknown significance (VUS). Genes of unknown significance aren't really talked about, because depending on position, variants within a gene can have different effects on the phenotype. It is the variant that is medically significant and gene-level annotations, while useful, are not determinative. --Mark viking (talk) 22:23, 12 January 2017 (UTC)
- Thanks for the link to Variants of unknown significance. Perhaps Gene of uncertain significance should be merged into it. Boghog (talk) 06:41, 13 January 2017 (UTC)
- @Maproom: I noticed that you took care of the winkler index issue at the helpdesk. VUS is the way I knew this concept, and I believe that is true for the vast majority of people who know about it. Why not just change the name of Gene of uncertain significance to "Variant of uncertain significance"? That way, both this article and the article Mark viking pointed us to can coexist. Since Variants of unknown significance is such a stub, let's have the longer gene article overwhelm it. BTW, to my eye the longer article isn't so bad. It needs work, but I'd be willng to help. DennisPietras (talk) 21:06, 15 January 2017 (UTC)
- @DennisPietras:: I am fairly sure I understand what the Winkler index article is about, so I checked some sources and went ahead with the move. I am less confident about the Variants of unknown significance material. It doesn't need any special power to move (rename) an article, you just need to be an autoconfirmed editor. If you feel confident that the move is justified, you can do it yourself: go to the cryptically named "More" menu at the top of the article, select its only item "Move", type in the new name, amd give your reason. Maproom (talk) 21:28, 15 January 2017 (UTC)
- @Maproom: I noticed that you took care of the winkler index issue at the helpdesk. VUS is the way I knew this concept, and I believe that is true for the vast majority of people who know about it. Why not just change the name of Gene of uncertain significance to "Variant of uncertain significance"? That way, both this article and the article Mark viking pointed us to can coexist. Since Variants of unknown significance is such a stub, let's have the longer gene article overwhelm it. BTW, to my eye the longer article isn't so bad. It needs work, but I'd be willng to help. DennisPietras (talk) 21:06, 15 January 2017 (UTC)
- Thanks for the link to Variants of unknown significance. Perhaps Gene of uncertain significance should be merged into it. Boghog (talk) 06:41, 13 January 2017 (UTC)
- Yes, but variant indicates an allele different from the reference genome, as in a VCF file. In clinical sequencing, we talk of variants and variants of unknown significance (VUS). Genes of unknown significance aren't really talked about, because depending on position, variants within a gene can have different effects on the phenotype. It is the variant that is medically significant and gene-level annotations, while useful, are not determinative. --Mark viking (talk) 22:23, 12 January 2017 (UTC)
- Aren't gene variants normally known as "alleles"? Maproom (talk) 22:03, 12 January 2017 (UTC)
done DennisPietras (talk) 04:14, 16 January 2017 (UTC)
- @Maproom, Boghog, Mark viking, and Aspro: I found a YouTube video made by a genetic counselor about VUS. She is speaking without a script, trying to educate people about VUS. To me, she made at least one mistake, but so be it. This topic is going to be a huge societal issue, and I think people need to hear it. It's here https://www.youtube.com/watch?v=cPFaAw4lSpU What I'd like to do is put some line of wikicode at the start of the article like [[Play_This_URL|as a right side thumbnail]]. Is that even possible? Would that be a violation of YouTube copyright? Do you think it is a good idea? If you think it's a good idea, possible, and not a violation, do any of you know what the real wikicode is? Thanks, DennisPietras (talk) 15:22, 17 January 2017 (UTC)
- There's more than one serious mistake in that video. Wikipedia ought be be able to do better. Maproom (talk) 16:51, 17 January 2017 (UTC)
- Yes, I agree that there are more mistakes, but would a poet who just got notice of a VUS notice them? The poet could benefit from actually hearing a voice repeating that it's not actionable and they need to keep in contact with their health care professional. Plus, I can't find a better shorter one! The ironic thing is that one of my former students is a genetic counselor, and she's let me down by not having such a video...8-( ..no I'm not going to contact her. — Preceding unsigned comment added by DennisPietras (talk • contribs) 18:57, 17 January 2017 (UTC)
- So there's such a thing as a VUS notice? That surprises me. Any two people (who aren't identical twins) must have hundreds of gene differences with some observable phenotypic effect, and thousands without. Maybe I've misunderstood something. Maproom (talk) 09:21, 18 January 2017 (UTC)
- @Maproom: Sit down for this: in your genome there are about 70 mutations that were not present in your parents' DNA! https://www.ncbi.nlm.nih.gov/pubmed/22345605 Off the top of my head, I believe I recall that each new whole genome sequence reveals something like 30 million basepair differences from the reference genome! Each of us have dozens of homozygous loss-of-function mutations! This is why it is so bloody difficult to make sense of variants that are found. If you have 25 minutes, watch, for example, https://www.youtube.com/watch?v=fC9rYghqUTo to get a feel for what medical professionals have to deal with! It may be the most amazing 25 minutes of yor life. With the cost of whole genome sequencing now at about $1,000 per individual, the flood of information is going to be overwhelming. Drinking from a full-open fire hydrant doesn't quite give an accurate feel. But the important question is whether any one of these changes has any phenotypic effect. That's why it is going to be so important to educate the public to understand that they don't need to freak out about VUS's. How did you feel when you just learned that you have dozens of homozygous LOF mutations? For me, it was comforting: I could blame my genes for all of my problems! But what about a 20 something planning on having a family? DennisPietras (talk) 13:53, 18 January 2017 (UTC)
- Ok, so its millions, not thousands. I can't say I'm surprised. But that makes it even more extraordinary that there should be such a thing as a VUS notice.
- @Maproom:I should have made my discussion clearer. For example, consider a woman with a first degree relative having breast cancer. The woman may decide to have her DNA sequenced to determine if she is at elevated risk for cancer from one of the know pathogenic BRCA gene mutations. She will get a lab report which will hopefully state that she does not have one of the known pathogenic mutations, and she will initially be overjoyed, but then go on to read the VUS notice (which I think/hope the lab is legaly required to provide) that she has a mutation in the gene that hasn't been linked to cancer development. The important thing is to educate the woman that she shouldn't focus on feeling "I've got a mutation in my BRCA1 gene" but to simply remember that and to keep her health care professionals aware of it and maybe keep herself up-to-date about the status of that variant, if she doesn't want to trust her health care professionals to keep track of the flood of VUS's that are a'comin'. DennisPietras (talk) 19:08, 18 January 2017 (UTC)
- Indeed. If she receives such a notice, everything possible should be done to encourage her to ignore it, and to convince her that it is the outcome of misguided bureaucracy. By "everything", I include explanations in WIkipedia articles. Maproom (talk) 23:20, 18 January 2017 (UTC)
- @Maproom:I should have made my discussion clearer. For example, consider a woman with a first degree relative having breast cancer. The woman may decide to have her DNA sequenced to determine if she is at elevated risk for cancer from one of the know pathogenic BRCA gene mutations. She will get a lab report which will hopefully state that she does not have one of the known pathogenic mutations, and she will initially be overjoyed, but then go on to read the VUS notice (which I think/hope the lab is legaly required to provide) that she has a mutation in the gene that hasn't been linked to cancer development. The important thing is to educate the woman that she shouldn't focus on feeling "I've got a mutation in my BRCA1 gene" but to simply remember that and to keep her health care professionals aware of it and maybe keep herself up-to-date about the status of that variant, if she doesn't want to trust her health care professionals to keep track of the flood of VUS's that are a'comin'. DennisPietras (talk) 19:08, 18 January 2017 (UTC)
- Ok, so its millions, not thousands. I can't say I'm surprised. But that makes it even more extraordinary that there should be such a thing as a VUS notice.
- @Maproom: Sit down for this: in your genome there are about 70 mutations that were not present in your parents' DNA! https://www.ncbi.nlm.nih.gov/pubmed/22345605 Off the top of my head, I believe I recall that each new whole genome sequence reveals something like 30 million basepair differences from the reference genome! Each of us have dozens of homozygous loss-of-function mutations! This is why it is so bloody difficult to make sense of variants that are found. If you have 25 minutes, watch, for example, https://www.youtube.com/watch?v=fC9rYghqUTo to get a feel for what medical professionals have to deal with! It may be the most amazing 25 minutes of yor life. With the cost of whole genome sequencing now at about $1,000 per individual, the flood of information is going to be overwhelming. Drinking from a full-open fire hydrant doesn't quite give an accurate feel. But the important question is whether any one of these changes has any phenotypic effect. That's why it is going to be so important to educate the public to understand that they don't need to freak out about VUS's. How did you feel when you just learned that you have dozens of homozygous LOF mutations? For me, it was comforting: I could blame my genes for all of my problems! But what about a 20 something planning on having a family? DennisPietras (talk) 13:53, 18 January 2017 (UTC)
- So there's such a thing as a VUS notice? That surprises me. Any two people (who aren't identical twins) must have hundreds of gene differences with some observable phenotypic effect, and thousands without. Maybe I've misunderstood something. Maproom (talk) 09:21, 18 January 2017 (UTC)
- Yes, I agree that there are more mistakes, but would a poet who just got notice of a VUS notice them? The poet could benefit from actually hearing a voice repeating that it's not actionable and they need to keep in contact with their health care professional. Plus, I can't find a better shorter one! The ironic thing is that one of my former students is a genetic counselor, and she's let me down by not having such a video...8-( ..no I'm not going to contact her. — Preceding unsigned comment added by DennisPietras (talk • contribs) 18:57, 17 January 2017 (UTC)
- There's more than one serious mistake in that video. Wikipedia ought be be able to do better. Maproom (talk) 16:51, 17 January 2017 (UTC)
WikiJournal of Medicine promotion

 |
The WikiJournal of Medicine is a free, peer reviewed academic journal which aims to provide a new mechanism for ensuring the accuracy of Wikipedia's biomedical content. We started it as a way of bridging the Wikipedia-academia gap.[1] It is also part of a WikiJournal User Group with other WikiJournals under development.[2] The journal is still starting out and not yet well known, so we are advertising ourselves to WikiProjects that might be interested. |
Engaging Wikipedians
- Original articles on topics that don't yet have a Wikipedia page, or only a stub/start
- Wikipedia articles that you are willing to see through external peer review (either solo or as in a group, process analogous to GA / FA review)
- Image articles, based around an important medical image or summary diagram
Engaging non-Wikipedians
We hope that an academic journal format may also encourage non-Wikipedians to contribute who would otherwise not. Therefore, please consider:
- Printing off the advertisement poster and distribute in tearooms & noticeboards at your place of work
- Emailing around the pdf through contact networks or mailing lists (suggested wording)
If you want to know more, we recently published an editorial describing how the journal developed.[3] Alternatively, check out the journal's About or Discussion pages.
- ^ Masukume, G; Kipersztok, L; Das, D; Shafee, T; Laurent, M; Heilman, J (November 2016). "Medical journals and Wikipedia: a global health matter". The Lancet Global Health. 4 (11): e791. doi:10.1016/S2214-109X(16)30254-6. PMID 27765289.
- ^ "Wikiversity Journal: A new user group". The Signpost. 2016-06-15.
- ^ Shafee, T; Das, D; Masukume, G; Häggström, M (2017). "WikiJournal of Medicine, the first Wikipedia-integrated academic journal". WikiJournal of Medicine. 4. doi:10.15347/wjm/2017.001.

Additionally, the WikiJournal of Science is just starting up under a similar model and looking for contributors. Firstly it is seeking editors to guide submissions through external academic peer review and format accepted articles. It is also encouraging submission of articles in the same format as Wiki.J.Med. If you're interested, please come and discuss the project on the journal's talk page, or the general discussion page for the WikiJournal User group.
T.Shafee(Evo&Evo)talk 10:33, 19 January 2017 (UTC)
Z-gene
@Akshaykatyura: Hi! I'm currently working my way from Z down to reclassify the unassigned articles. The Z-gene article https://en.wikipedia.org/wiki/Z-gene is a stub about the lac z gene, which should be converted to a redirect to the lac operon, IMHO. Any objections? DennisPietras (talk) 03:58, 23 January 2017 (UTC)
- Agreed. Good catch. Ajpolino (talk) 04:59, 23 January 2017 (UTC)
- A disambiguation page would be better, as "Z gene" could also refer to Protein Z or to an unrelated gene involved in avian sex determination ([2]). Adrian J. Hunter(talk•contribs) 05:40, 23 January 2017 (UTC)
- Also if someone who knew anything about the lac operon were to type "Z-gene" into the search box, they'd presumably be looking for Beta-galactosidase rather than lac operon. Adrian J. Hunter(talk•contribs) 05:44, 23 January 2017 (UTC)
- I boldly converted it to a disambig. Adrian J. Hunter(talk•contribs) 06:46, 23 January 2017 (UTC)
- @Adrian J. Hunter:
 Thank you DennisPietras (talk) 02:19, 24 January 2017 (UTC)
Thank you DennisPietras (talk) 02:19, 24 January 2017 (UTC)
- @Adrian J. Hunter:
- I boldly converted it to a disambig. Adrian J. Hunter(talk•contribs) 06:46, 23 January 2017 (UTC)
PIC and mediator
Hi! I've noticed that there are separate Transcription preinitiation complex and Mediator (coactivator) pages. I think that based on a 2015 review, http://www.nature.com/nrm/journal/v16/n3/box/nrm3951_BX3.html the pages should be merged. Comments? Thanks, DennisPietras (talk) 03:30, 25 January 2017 (UTC)
- It may not need to be fully merged. We have separate pages for TFIIA etc. The mediator complex seems to be a well-defined group of proteins, but absolutely needs to be mentioned as a key component of the PIC on the Transcription preinitiation complex page. T.Shafee(Evo&Evo)talk 03:36, 25 January 2017 (UTC)
- I agree with Thomas. The review article states that the mediator coactivator complex has roles in addition to being a component of the preinitiation complex. Hence I think it is better to keep the two articles separate while mentioning that mediator is a component of PIC. Boghog (talk) 04:29, 25 January 2017 (UTC)
the forlorn 19
There were 19 articles listed as stub quality and NA importance. Almost all of them were redirect pages, and I decided to be bold and just remove the genetics project banner from those pages. In the "can you believe it" category, one of the pages that was redirected to was site directed mutagenesis, which didn't have a mention of CRISPR on it! So much to do, so little time. The 19 are still there as I write this, because the bot doesn't work in real time. DennisPietras (talk) 19:15, 18 January 2017 (UTC)
- I also hopefully took care of the forlorn 12 rated start quality and NA importance. — Preceding unsigned comment added by DennisPietras (talk • contribs) 19:40, 18 January 2017 (UTC)
- Good work. I've been working through the '???' importance rating category. T.Shafee(Evo&Evo)talk 23:33, 18 January 2017 (UTC)
- @DennisPietras: I can absolutely recommend the "Rater" plugin by Kephir. Just go to your Special:MyPage/common.js and add the text
importScript('User:Kephir/gadgets/rater.js'); // [[User:Kephir/gadgets/rater]]. It allows you to edit the class and importance of a page from the Article (without having to switch back and forth to the talk page), and auto-fills an edit summary describing the change so that it's easier to see when people look back through the history tab. T.Shafee(Evo&Evo)talk 11:14, 25 January 2017 (UTC)- @Evolution and evolvability: Remarkably (to me - I don't know how you folks get to know all these things) I actually did get "rater" to appear to the left of "more" on my page. I click on it, and a box opens that appears to indicate that it is a beta version, but, in any case I'm lost about how to actually use it. This may beyond my comprehension, and I'm fairly comfortable doing the ratings the slow way, because I get to see what's up on the talk page. Thanks, DennisPietras (talk) 00:44, 26 January 2017 (UTC)
- @Evolution and evolvability: DUH Now I see how to use it wen I'm on the page I want to rate!
 Thank you DennisPietras (talk) 00:48, 26 January 2017 (UTC)
Thank you DennisPietras (talk) 00:48, 26 January 2017 (UTC)
- @Evolution and evolvability: DUH Now I see how to use it wen I'm on the page I want to rate!
- @Evolution and evolvability: Remarkably (to me - I don't know how you folks get to know all these things) I actually did get "rater" to appear to the left of "more" on my page. I click on it, and a box opens that appears to indicate that it is a beta version, but, in any case I'm lost about how to actually use it. This may beyond my comprehension, and I'm fairly comfortable doing the ratings the slow way, because I get to see what's up on the talk page. Thanks, DennisPietras (talk) 00:44, 26 January 2017 (UTC)
- @DennisPietras: I can absolutely recommend the "Rater" plugin by Kephir. Just go to your Special:MyPage/common.js and add the text
- Good work. I've been working through the '???' importance rating category. T.Shafee(Evo&Evo)talk 23:33, 18 January 2017 (UTC)
@Evolution and evolvability: Much to my delight, I find that rater gives me the option to rate pages importance=bottom. I've been using that for books, etc. However, when I go back to look at the list you generated for me, the "bottom" articles are still listed. I'm going to go ahead and keep using that bottom rating at least until the "bot" (or whatever) problem we've been talking about is fixed. But, I'm wondering if the table "Genetics articles by quality and importance" needs to be modified to have a "bottom" row? DennisPietras (talk) 19:48, 26 January 2017 (UTC)
Is there a way to get a list of genetics article sorted by their average page views?
Hi! I know where/how to get page view statistics for individual articles, but that would be frustrating, to say the least, for all artcles. Is there a way to produce a list of all articles in the genetics project sorted by page views, hopefully with number of page views listed next to the articles? I have a vague feeling that a SQL search could generate such a list, but my knowledge of SQL ended in the 1990's. Thanks, DennisPietras (talk) 17:30, 2 February 2017 (UTC)
- There is the script at User:Dsimic/Traffic stats calculation that given a list of articles, produces page views. But I haven't tried it myself. --Mark viking (talk) 20:01, 2 February 2017 (UTC)
- I just looked at that script, and it is waaaayyyy over my head. What I was really hoping for by starting this topic was to nudge Thomas into doing it for us!
 DennisPietras (talk) 22:35, 2 February 2017 (UTC)
DennisPietras (talk) 22:35, 2 February 2017 (UTC)
- I just looked at that script, and it is waaaayyyy over my head. What I was really hoping for by starting this topic was to nudge Thomas into doing it for us!
- The WP:GEN assessment page has a list of the top 500, but that's still pretty minimal dataset. What you probably want is the MASSVIEWS wikimedia foundation tool that allows you to view the stats for all articles in any category. E.g. Here is the one for everything in Category:WikiProject Genetics articles (don't forget to check the "use subject page" tickbox). You could similarly do one for just articles in specific imporance/class categories (e.g. Category:C-Class_Genetics_articles or Category:Top-importance_Genetics_articles). T.Shafee(Evo&Evo)talk 01:02, 3 February 2017 (UTC)
- Wow! Right under my nose, so to speak. I'm going to have to stop using the newbie excuse soon... Thanks Thomas! DennisPietras (talk) 03:38, 3 February 2017 (UTC)
- Absolutely no problem! There can be a lot to get used to on Wikipedia. I'm still learning things after years here. T.Shafee(Evo&Evo)talk 12:04, 3 February 2017 (UTC)
- Wow! Right under my nose, so to speak. I'm going to have to stop using the newbie excuse soon... Thanks Thomas! DennisPietras (talk) 03:38, 3 February 2017 (UTC)
- The WP:GEN assessment page has a list of the top 500, but that's still pretty minimal dataset. What you probably want is the MASSVIEWS wikimedia foundation tool that allows you to view the stats for all articles in any category. E.g. Here is the one for everything in Category:WikiProject Genetics articles (don't forget to check the "use subject page" tickbox). You could similarly do one for just articles in specific imporance/class categories (e.g. Category:C-Class_Genetics_articles or Category:Top-importance_Genetics_articles). T.Shafee(Evo&Evo)talk 01:02, 3 February 2017 (UTC)
proposed change to "to do list"
Hi! Currently, the task is:
Please help to assess the importance and quality of these unassessed articles
which is almost done. So, looking ahead, I suggest that the to do list read:
"There are currently 7 articles rated as stub-class and high-importance. They are, with the average daily number of page views listed after them
- TATA box 355
- coding strand 235
- coding region 174
- germline mutation 102
- Pribnow box 81
- gene delivery 28
- Nuclear gene 18
If you are interested in editing, please help improve these pages! Even if you are new to editing, don't worry about making mistakes. People will be watching and helping as needed."
Comments? Objections? DennisPietras (talk) 15:50, 3 February 2017 (UTC)
I guess this proves that I'm a masochist - in the asexual sense
So, the page https://en.wikipedia.org/wiki/Wikipedia:WikiProject_Genetics/Assessment gives guidance about how to rate the importance of articles. I wonder if we should let the community of wikipedia readers rate the importance of articles. That is, if a page averages more than 17 page views/day it becomes rated at least high importance. I set the level at 17 so that nuclear gene would stay at high importance. Heterogeneous ribonucleoprotein particle is a stub currently rated at mid importance, but it has 43 views/day. By my suggestion, it would become high importance, even though I don't think it qualifies by the current guideline of "High school students may have some familiarity, or else early undergraduate students should be very familiar with the subject." Believe me, I realize that this reclassification would be a lot of work, but the one person who has adopted gene delivery for a school project has re-energized me for the task. Comments? DennisPietras (talk) 04:19, 10 February 2017 (UTC)
- @DennisPietras: Sorry to be a dampener on this, but I think that the importance ranking and reader traffic are separate statistics. Firstly, traffic can stats fluctuate a lot over time depending on what's in the news, and I don't think it's useful for the importance rating to keep up with that. Also, importance can be different to different WikiProjects, e.g. Axolotl is high-importance to WikiProject Amphibians and Reptiles, but the genetics components of that page are only of low importance to WP:GEN. Finally, I think it's also useful to keep more-or-less in line with other WikiProjects. I don't mean to be overly-negative! I think that the current ratings guidelines are reasonably sensible, even if their implementation is still quite variable and subjective. I totally agree with your move of CRISPR to "High" though! T.Shafee(Evo&Evo)talk 11:25, 10 February 2017 (UTC)
- I'm not sure Nuclear gene should be high priority. It's really just a term that needs to be defined... The relevant info mostly belongs at Gene, Mitochondria, etc. 17 views per day is tiny compared to, say, Genome or Dominance (genetics), each at ~900 views per day. I think setting a 17 view per day cutoff would lead to a huge number of high-importance articles, which would defeat the purpose of the rating. Adrian J. Hunter(talk•contribs) 12:10, 10 February 2017 (UTC)
Mediator (coactivator)
Hi again folks. I'm starting to gather up the references and courage to tackle improvements to Mediator (coactivator), which gets 63 views/day. I have 2 separate concerns.
- Does anybody object to me moving the article to "Mediator complex (molecular biology)"?
- The article has had very few recent edits. Boghog is by far the most recent human editor (presuming that her/his username isn't reflective of his/her species identity
 ). What I would prefer to do is copy the code of the page into my sandbox, work on it there, and then transfer the whole thing back into the mediator page when I'm done. That's what I did at https://en.wikipedia.org/wiki/User:DennisPietras/sandbox3 before transfering it into the Proximity ligation assay page, and I liked that much more than tryiing to work on an article when it is "live". Is that the way major edits are usually done? I wouldn't suggest it, except that Boghog now knows my intentions and there is little other editorial interest. Should I put an "under construction" sign on mediator even though I'm working on it in my sandbox? Any suggestions? Thanks, DennisPietras (talk) 15:06, 10 February 2017 (UTC)
). What I would prefer to do is copy the code of the page into my sandbox, work on it there, and then transfer the whole thing back into the mediator page when I'm done. That's what I did at https://en.wikipedia.org/wiki/User:DennisPietras/sandbox3 before transfering it into the Proximity ligation assay page, and I liked that much more than tryiing to work on an article when it is "live". Is that the way major edits are usually done? I wouldn't suggest it, except that Boghog now knows my intentions and there is little other editorial interest. Should I put an "under construction" sign on mediator even though I'm working on it in my sandbox? Any suggestions? Thanks, DennisPietras (talk) 15:06, 10 February 2017 (UTC)
- Since there is another article called Endogenous mediator, renaming Mediator (coactivator) as Mediator complex (molecular biology) would not be appropriate as it would be too general. Even if these two articles were merged, I still not sure the rename would be appropriate. Wikipedia generally does not disambiguate article names based on the field. For example, the disambiguations for Mercury are Mercury (element), not Mercury (chemistry) and Mercury (planet), not Mercury (astronomy). Generally there is no need to work on an article in your sandbox and in fact reasons not to, especially if the article is actively being edited by others. Also if you completely restructure an article in your sandbox and then apply those changes in one edit to the live article, others might object. It is better to make smaller edits one at a time and see how others react. While there is no requirement to do so, if you plan to completely restructure an article, it would be prudent to briefly outline the changes you plan to make on the article's talk page before your make those changes to see what others think. Boghog (talk) 19:58, 10 February 2017 (UTC)
@Fnielsen: et al. Does anybody object to me merging Asp... into MC1R and then making Asp... into a redirect to MC1R? DennisPietras (talk) 02:31, 5 February 2017 (UTC)
- Haha, definite merge. I'm not convinced that ever warranted its own article. V600E shoudl also be merged into BRAF and C957T could even be merged into DRD2. In fact, the only article on a specific gene mutation that I've ever seen done well is ΔF508, which has a lot of information that usefully expands on the Cystic fibrosis article. T.Shafee(Evo&Evo)talk 00:53, 6 February 2017 (UTC)
- You can merge it, but let the Wikidata item stay. Another (half) example: Factor V Leiden is written as a genetic disorder, but has the "Infobox single nucleotide polymorphism". — fnielsen (talk) 15:04, 7 February 2017 (UTC)
- @Fnielsen and Evolution and evolvability:Wow, I've opened a nice can of worms! I'm learning so many more things about MCR1 that I'm intending to make major additions to that article. So, now what I want to do specifically with Asp294His is to just move it to "MC1R Asp294His". Any objections? Thanks, DennisPietras (talk) 19:24, 7 February 2017 (UTC)
- @DennisPietras: Given that it's so short, I would have through ti could just be a section of the Melanocortin_1_receptor page, perhaps in a section titled "single nucleotide polymorphisms". However if you are happy to expand it sufficiently to justify its existence, I'd go for Melanocortin_1_receptor D294H as a title, to include the protein's full name, and use the shorter notation for the mutation. However, as you've noticed, we've not really any strong precedents for these sorts of SNP articles, since there are so few, and they're a tiny fraction of the possible notable SNPs. T.Shafee(Evo&Evo)talk 03:26, 8 February 2017 (UTC)
- @Evolution and evolvability:I think you misunderstood. I left the snp article untouched, and just finished a distressingly long process of beefing up the MC1R article. Off to bed. DennisPietras (talk) 06:27, 8 February 2017 (UTC)
- @Fnielsen and Evolution and evolvability: I just moved it to Melanocortin 1 receptor Asp294His DennisPietras (talk) 03:39, 10 February 2017 (UTC)
- @Evolution and evolvability:I think you misunderstood. I left the snp article untouched, and just finished a distressingly long process of beefing up the MC1R article. Off to bed. DennisPietras (talk) 06:27, 8 February 2017 (UTC)
- @DennisPietras: Given that it's so short, I would have through ti could just be a section of the Melanocortin_1_receptor page, perhaps in a section titled "single nucleotide polymorphisms". However if you are happy to expand it sufficiently to justify its existence, I'd go for Melanocortin_1_receptor D294H as a title, to include the protein's full name, and use the shorter notation for the mutation. However, as you've noticed, we've not really any strong precedents for these sorts of SNP articles, since there are so few, and they're a tiny fraction of the possible notable SNPs. T.Shafee(Evo&Evo)talk 03:26, 8 February 2017 (UTC)
- @Fnielsen and Evolution and evolvability:Wow, I've opened a nice can of worms! I'm learning so many more things about MCR1 that I'm intending to make major additions to that article. So, now what I want to do specifically with Asp294His is to just move it to "MC1R Asp294His". Any objections? Thanks, DennisPietras (talk) 19:24, 7 February 2017 (UTC)
- You can merge it, but let the Wikidata item stay. Another (half) example: Factor V Leiden is written as a genetic disorder, but has the "Infobox single nucleotide polymorphism". — fnielsen (talk) 15:04, 7 February 2017 (UTC)
With a general remark: It seems that science beginning perhaps in the 1990s had great optimism regarding individual SNPs to explain a variety of behaviour and diseases. I have not so much followed the topic for the past years but it seems to me that "SNP-blaming" is not so much in fashion anymore. Later meta-analysis studies may have squashed hopes. Perhaps it is me that is not up to date? Should articles such as Melanocortin 1 receptor Asp294His be regarded as a fad of the 00s? — fnielsen (talk) 08:49, 10 February 2017 (UTC)
- @Fnielsen: A good point. There are certainly a few SNPs that have been shown to have large enough effects even on their own, so I think it is conceivable that there could be sufficiently notable SNPs to justify their own article. However I don't know enough about the topic to know if those actually match up with the handful of SNP articles that currently exist. In general I favour merging into parent gene articles since I suspect that otherwise readers don't find them anyway. T.Shafee(Evo&Evo)talk 10:47, 10 February 2017 (UTC)
- @Fnielsen and Evolution and evolvability: et al. Judging from the number of GWAS studies, I don't think snips are a fad, but I do agree with Thomas that they should generally be merged into the parent gene article. DennisPietras (talk) 21:35, 10 February 2017 (UTC)
genetics of cancer redirect to oncogenomics
@FourViolas: I've disccovered that there is a page "genetics of cancer" which is only a "start" quality article. Meanwhile, "Oncogenomics" is B quality. I propose making genetics of cancer a simple redirect to oncogenomics. Comments, especially from FourViolas, who appears to be the only one recently watching c.o.g? Thanks, DennisPietras (talk) 20:40, 30 January 2017 (UTC)
- No objections on the genetics of cancer talk page for more than a week, so I made the redirect. DennisPietras (talk) 02:00, 14 February 2017 (UTC)
Subunit composition section and table
Hi again. The mediator Subunit composition section begins "The Mediator complex is composed of up to at least 31 subunits in all eukaryotes studied:" which is only true because the editor included "up to" as in advertisements that state up to 75% off.![]() The main question I have is this: Do we really want to list all of the subunits and include the table? My thought is to put links to subunits that have wp articles in a "see also" section and eliminate the table and "subunit composition" section entirely. I haven't checked completely (nor do I intend to) but the table was added in 2012 and I have to believe it is incomplete. Thanks for any comments. DennisPietras (talk) 21:02, 14 February 2017 (UTC)
The main question I have is this: Do we really want to list all of the subunits and include the table? My thought is to put links to subunits that have wp articles in a "see also" section and eliminate the table and "subunit composition" section entirely. I haven't checked completely (nor do I intend to) but the table was added in 2012 and I have to believe it is incomplete. Thanks for any comments. DennisPietras (talk) 21:02, 14 February 2017 (UTC)
- The human subunits are already listed in the subunit composition sectdion. The table is a cross-species comparison. Boghog (talk) 07:19, 15 February 2017 (UTC)
Hi again folks. Transcription factor gets about 700 views/day. General tf about 100. Is there any traction out there for merging the 2 articles? I came across those 2 while updating mediator (which is far from finished), and then clearly preinitiation complex is next for me. DennisPietras (talk) 02:42, 14 February 2017 (UTC)
- The term "general transcription factor" is frequently used in the literature and hence is independently notable and consequently deserves its own article. I see no advantage of merging these two articles. Boghog (talk) 06:35, 15 February 2017 (UTC)
- I agree with Boghog. They're distinct enough topics that they warrant being separate. Also the Transcription factor article is already pretty long. T.Shafee(Evo&Evo)talk 23:03, 15 February 2017 (UTC)
Mediator explanatory footnotes
Hi again folks. I'd appreciate feedback on how I'm using enf's in Mediator (coactivator), particularly how I used it in enf d to direct folks to be able to see a reasonably-sized version of a copyrighted image that I can't put on the commons. Actually, I'd most appreciate some "way to go's", but if I'm violating wp community standards, it's better to know now. Thanks, DennisPietras (talk) 20:48, 14 February 2017 (UTC)
- Thanks for your additions. I don't have any strong feeling about footnotes except that they make the article more complicated to read and to maintain. One thing to keep in mind is that graphics are intended to support the text, not the other way around. Also it is assumed that the figures are simplifications of reality and it is not necessary to dwell on their shortcomings. Boghog (talk) 07:15, 15 February 2017 (UTC)
- Good work so far DennisPietras. My recommendations are to make sure the lead section only summarises info that's in the rest of the article. Currently the lead seems a bit long. Moving the majority of the information into its own section will also allow more space for the images. As for the footnotes, I think they end up rarely being read, since readers seldom distinguish them from references so don't think to click on most. However there's certainly no harm in them. An alternative strategy for the copyrighted image is to email the author of the paper and ask if they have another version that isn't copyrighted or is under a creative commons license. I've found that people are often very keen to contribute their images once they know what they're being used for. T.Shafee(Evo&Evo)talk 23:01, 15 February 2017 (UTC)
- Thanks to both of you. Re: the lead length: it will appear short when I'm done! The reason I put several images in so far is to emphasize the flexibility in structure and associations. That can't be done with just one image, and, yes, I was reluctant to put one on the left, as it is discouarged by wp, but for the foreseeable future I think it will stay there OK. Re: footnotes and readability: I expect that the vast majority of readers won't look at the ref's or efn's, which is why I think putting material in the efn's makes the article more readable. Then, if the article tickles the reader's fancy, they can go back and look at the efn's and/or ref's. Re: image contribution: if I got permission to use more, then I'd have to find a place to put them!!!!!!! If a reader fancies, they can find the images from the info I've given them.
 Thank you DennisPietras (talk) 01:20, 16 February 2017 (UTC)
Thank you DennisPietras (talk) 01:20, 16 February 2017 (UTC)
- Thanks to both of you. Re: the lead length: it will appear short when I'm done! The reason I put several images in so far is to emphasize the flexibility in structure and associations. That can't be done with just one image, and, yes, I was reluctant to put one on the left, as it is discouarged by wp, but for the foreseeable future I think it will stay there OK. Re: footnotes and readability: I expect that the vast majority of readers won't look at the ref's or efn's, which is why I think putting material in the efn's makes the article more readable. Then, if the article tickles the reader's fancy, they can go back and look at the efn's and/or ref's. Re: image contribution: if I got permission to use more, then I'd have to find a place to put them!!!!!!! If a reader fancies, they can find the images from the info I've given them.
- Good work so far DennisPietras. My recommendations are to make sure the lead section only summarises info that's in the rest of the article. Currently the lead seems a bit long. Moving the majority of the information into its own section will also allow more space for the images. As for the footnotes, I think they end up rarely being read, since readers seldom distinguish them from references so don't think to click on most. However there's certainly no harm in them. An alternative strategy for the copyrighted image is to email the author of the paper and ask if they have another version that isn't copyrighted or is under a creative commons license. I've found that people are often very keen to contribute their images once they know what they're being used for. T.Shafee(Evo&Evo)talk 23:01, 15 February 2017 (UTC)
Am I alone in my frustration?
Hi again folks. It seems like everytime I set out to update a genetics article it leads to 2 more that desperately need upgrading! It's like playing Whac-A-Mole for real! sigh. Is this a common experience? Can anybody suggest an alternative treatment to the one I use - eating party peanuts until I forget that I'm frustrated? Thanks, DennisPietras (talk) 20:19, 21 February 2017 (UTC)
- yep WP is very uneven -- great in spots, terrible in spots, and mediocre in many; it is a direct result of this being a crowd-sourced beast. There is a ton of work to do, always. You just need to pace yourself and make sure the work you do is very high quality (great sources, neutrally summarized, and overall WEIGHT correct) so that what you actually spend time on sticks, and you can move on to the next problematic article. Jytdog (talk) 20:34, 21 February 2017 (UTC)
- I don't think you are alone. It is natural to want to see a field one cares about having lots of solid articles chock full of relevant reliable information, and it is disappointing when most articles fall short. But WP is very much a work in progress--it is a construction site with lots of partially finished articles, some barely started. Rather than feeling frustration, though, I like to see it as a great opportunity to contribute and improve the encyclopedia. With so many articles having potential for improvement, it can be tough to choose which to work on. Jytdog's advice is good here--pick an article in which interests you the most and for which you have the knowledge and resources to add solid content. --Mark viking (talk) 21:02, 21 February 2017 (UTC)
- I agree with the comments above. I suspect we've found ourselves lost down editing rabbit holes at some point! I tend to try to set specific and achievable goals (e.g. update a specific article section or make a new image), otherwise it's easy to focus on what's left to do, rather than receive the endorphins due for what you've already done. It think that that is why people often have little trophy cabinets of things they're proud of on their userpages. Every month I leave the world in a slightly better state than I found it. Then again, sometimes I actually enjoy just following a chain of wikilinks, spending 2-5 mins on each article's wording, or reference formatting (WP:WikiGnome), since it requires less brain. Another thing that I find helpful is to occasionally read news about Wiki stuff (e.g. The Signpost and the WMF blog). In part, it just takes practice to find what works best for you! T.Shafee(Evo&Evo)talk 00:56, 22 February 2017 (UTC)
 Thank you to all! I'm feelng better today. DennisPietras (talk) 03:40, 23 February 2017 (UTC)
Thank you to all! I'm feelng better today. DennisPietras (talk) 03:40, 23 February 2017 (UTC)
Opinions requested
Opinions are requested at Talk:Expanded genetic code#mouse code about whether certain content is an acceptable use of primary sources. Thanks, Adrian J. Hunter(talk•contribs) 06:10, 13 March 2017 (UTC)
revisions to pioneer factor
Hi folks! I've been working on revisions to the pioneer factor page on my sandbox, and I am done, IMHO, with the text as I envision it as of 3/14/17. Please review and/or edit User:DennisPietras/sandbox5 if you have time and let's try to reach a consensus on what the wp page should be, before transfering the sandbox to the mainspace. Thanks, DennisPietras (talk) 00:17, 15 March 2017 (UTC)
Proposal of using full size image for RNA expression pattern
Hello WikiProject Genetics! In infoboxes of gene/protein articles, there are small thumbnail images of RNA expression pattern. I proposed switching these small thumbnail to full size image, because Wikipedia/Mediawiki image system was changed. Since image data are recalled from Wikidata, I post the proposal at Wikidata page (wikidata:Property talk:P692#How about using full size image instead of small thumbnail?). I would like to get your thoughts on that. Thank you. --Was a bee (talk) 06:32, 18 March 2017 (UTC)
Expressomics redirected to Transcriptome
There is a page expressomics that was created by Jongbak in 2007. Transcriptomics is the current term, but transcriptomics is a simple redirect to transcriptome. I propose that expressomics become a simple redirect to transcriptome. Any objections? DennisPietras (talk) 21:39, 1 February 2017 (UTC)
- A merge would be better as each article has unique content, and per WP:PRESERVE, we usually try to preserve verifiable content. For instance, the expressomics article has a section on assay techniques mostly missing from transcriptome--indeed the scope section in transcriptome is completely unsourced.. --Mark viking (talk) 19:48, 2 February 2017 (UTC)
- Yes, of course, merge first. Remember, I'm a newbie...I wonder how much longer I'll be able to use that excuse for my brain cramps...
 DennisPietras (talk) 22:28, 2 February 2017 (UTC)
DennisPietras (talk) 22:28, 2 February 2017 (UTC)
- Yes, of course, merge first. Remember, I'm a newbie...I wonder how much longer I'll be able to use that excuse for my brain cramps...
Good idea. I added this proposal to each page. Discuss Talk:Transcriptome#Merge Expressomics page into Transcriptome — Preceding unsigned comment added by MangoldOrganizer (talk • contribs) 03:11, 25 March 2017 (UTC)
Extension of 'Topic Page' review articles from PLOS Computational Biology to PLOS Genetics

The journal group PLOS is extending its 'Topic Page' review format that was spearheaded by PLOS Computational Biology to also include PLOS Genetics. In this format, accepted articles are dual-published both in the journal, and as Wikipedia pages (see Wikipedia category).
Suitable topics must either currently lack a Wikipedia page, or have only stub/start class contents. If you you would like to submit such a review article, see these guidelines. If you have any recommendations for topics to be commissioned, feel free to let any of the involved editors know: T Shafee (PLOS Gen), D Mietchen (PLOS Comp Biol).
T.Shafee(Evo&Evo)talk 12:26, 25 March 2017 (UTC)
- Exciting! One small bug: the guideline linked to only mentions stub class articles as being suitable. I'd be in favor of including start class articles as suitable as well. --Mark viking (talk) 12:53, 25 March 2017 (UTC)
- Well spotted, I've corrected it on the PLOS wiki page. T.Shafee(Evo&Evo)talk 00:37, 26 March 2017 (UTC)
- A blog post on the matter is now out. Re the stubs vs. start discussion above, this was intentional when we launched it for Comp Bio but I agree that nothing has come up since that would prevent extension to start class articles. -- Daniel Mietchen (talk) 11:27, 13 April 2017 (UTC)
- Well spotted, I've corrected it on the PLOS wiki page. T.Shafee(Evo&Evo)talk 00:37, 26 March 2017 (UTC)
Russian Domesticated Red Fox
Greetings to the Genetics project members! Kindly note my assessment in Talk:Russian Domesticated Red Fox#Popular book published in March 2017; further editing needed. I found the Russian Domesticated Red Fox page problematic (see my two edits), though I lack the scientific background to properly improve it. Is it correct that the more extensive description of the experimental project is found on the biographical page of the lead scientist, Dmitry Belyayev? Your help is most welcome, and I look forward to learning from others' further edits. -- Deborahjay (talk) 12:16, 7 May 2017 (UTC)
Popular pages report
We – Community Tech – are happy to announce that the Popular pages bot is back up-and-running (after a one year hiatus)! You're receiving this message because your WikiProject or task force is signed up to receive the popular pages report. Every month, Community Tech bot will post at Wikipedia:WikiProject Molecular Biology/Genetics/Archive 3/Popular pages with a list of the most-viewed pages over the previous month that are within the scope of WikiProject Molecular Biology.
We've made some enhancements to the original report. Here's what's new:
- The pageview data includes both desktop and mobile data.
- The report will include a link to the pageviews tool for each article, to dig deeper into any surprises or anomalies.
- The report will include the total pageviews for the entire project (including redirects).
We're grateful to Mr.Z-man for his original Mr.Z-bot, and we wish his bot a happy robot retirement. Just as before, we hope the popular pages reports will aid you in understanding the reach of WikiProject Molecular Biology, and what articles may be deserving of more attention. If you have any questions or concerns please contact us at m:User talk:Community Tech bot.
Warm regards, the Community Tech Team 17:16, 17 May 2017 (UTC)
Figures at Recombinase-mediated cassette exchange
Hello WikiProject Genetics. I am here regarding the article Recombinase-mediated cassette exchange, and I'm hoping you can resolve an issue I'm seeing. A friendly reader contacted the Volunteer Response Team the other day (VRTS ticket # 2017071510006261) and pointed out that the two figures in that article are identical and that the caption to Figure 1 doesn't seem to make sense. I took a look and, indeed, the two figures, which have different captions that refer to different things, are both copies of File:RMCE2.PNG.
Now, I looked in the history of the article, and it appears in 2013, the article had another image, the now-deleted image File:RMCE.PNG. This file was deleted on the basis that the original uploader had uploaded an "updated" image, but it appears the "updated" image might have been mistaken to be File:RMCE2.PNG, an unrelated image. Perhaps File:RMCE.PNG should be undeleted and restored as Figure 1 in the article? But why was it removed in the first place? If necessary, I can provisionally undelete the image so that it can be reviewed. Mz7 (talk) 04:11, 17 July 2017 (UTC)
- Okay, I contacted the administrator who originally deleted File:RMCE.PNG, and he has now restored it, and I've restored it into the article again. Hopefully this will resolve the reader's concern. Let me know if you guys spot anything. Mz7 (talk) 07:22, 19 July 2017 (UTC)
Genetics and archaeogenetics of Central Asia
If you search for such subjects you are redirected / sent to Genetics and archaeogenetics of South Asia. Why not make a seperate article? I could start a stub article, but I worry there may be reason for the lack of such a page. — Preceding unsigned comment added by Darokrithia (talk • contribs) 19:27, 7 August 2017 (UTC)
- @Darokrithia: I think the redirect is probably there just because of lack of content. It's not really my area, but my suggestion would be to replace the redirect at Genetics and archaeogenetics of Central Asia with a stub on the specific topic and see if anyone reverts it. Remember to put a "See also" section pointing to the South Asia version of the article. You might also want to check with WikiProject Human Genetic History in case they have any ideas. T.Shafee(Evo&Evo)talk 01:40, 8 August 2017 (UTC)
Gene location column added to infobox gene
New column which shows gene location is added to infobox gene. Feedback welcome at Module talk:Infobox gene#Gene location column added. Thanks! --Was a bee (talk) 11:43, 18 August 2017 (UTC)
SNPs
Hi! I found that Wikipedia has almost no articles about SNPs (Single-nucleotide polymorphism). Although there are over 140 thousands SNPs in SNPedia, I found only 22 articles in Wikipedia. I categorized them according to their positions on human chromosomes e.g. Category:SNPs on chromosome 1, Category:SNPs on chromosome 11. I think more work should be done on expanding articles about this topic. Here is a list of over 140,000 SNPs, if anyone is interested in creating new SNPs articles. I also suggest starting a special Wikiproject for SNPs expansion. --Brainist (talk) 20:39, 20 November 2017 (UTC)
- Hi Brainist, I'm curious to know what others think, but I'm not sure Wikipedia is the best project for this sort of information. Most of these articles seem to be of borderline notability. Wikidata or some dedicated project would be better. There might be a few notable SNPs like rs6265, but even in that case the information would be better contextualised in the article on the parent gene. Btw not everything in dbSNP is really a SNP – eg Rs28363170 is not. Adrian J. Hunter(talk•contribs) 23:39, 20 November 2017 (UTC)
- I would have to agree on this. It would be a really high bar for a SNP to be notable, but by then the underlying gene should already be known. An article on a SNP would definitely be an exception rather than an expectation. Kingofaces43 (talk) 03:27, 21 November 2017 (UTC)
- Wikidata could be a perfect unifying database for this sort of thing (Andrew Su may have some ideas via their experience with gene wiki). There are very few SNPs that have sufficient info to make an article about. I think that the majority would be better merged into the article on their parent gene. Although there will certainly be a few that are sufficiently notable in their own right (e.g. perhaps the SNP that causes sickle cell anaemia β-globin E7V), but I think they should only get split into their own page if the parent article gets too overwhelmed by content. T.Shafee(Evo&Evo)talk 09:07, 21 November 2017 (UTC)
- I agree. If anyone is interested in the wikidata angle, I suggest posting here. Best, Andrew Su (talk) 16:27, 21 November 2017 (UTC)
- Chiming in from WikiProject Computational Biology, I agree with the others here: Wikidata would be a better solution for as the majority of SNPs won't be generally notable. I'd definitely be interested in helping out if there's a consensus to migrate info from SNPedia to Wikidata. Thanks, Amkilpatrick (talk) 09:22, 22 November 2017 (UTC)
- This isn't a terribly active WikiProject, Amkilpatrick. What you see right here is about as much of a consensus as you're ever likely to get. Adrian J. Hunter(talk•contribs) 23:30, 22 November 2017 (UTC)
- Wikidata could be a perfect unifying database for this sort of thing (Andrew Su may have some ideas via their experience with gene wiki). There are very few SNPs that have sufficient info to make an article about. I think that the majority would be better merged into the article on their parent gene. Although there will certainly be a few that are sufficiently notable in their own right (e.g. perhaps the SNP that causes sickle cell anaemia β-globin E7V), but I think they should only get split into their own page if the parent article gets too overwhelmed by content. T.Shafee(Evo&Evo)talk 09:07, 21 November 2017 (UTC)
- I would have to agree on this. It would be a really high bar for a SNP to be notable, but by then the underlying gene should already be known. An article on a SNP would definitely be an exception rather than an expectation. Kingofaces43 (talk) 03:27, 21 November 2017 (UTC)
Genetic Engineering Taskforce
This is a controversial area on Wikipedia as some watching here may know. I have been editing it on and off for seven years and now that the dust has settled from the ARBCOM case I am hoping to get the content in this area organised and improved. I have just got Genetic Engineering to good article status so it is possible. I am happy to plod on by myself, but it is a big and complicated area so any help would be appreciated. There are also a lot of comments to talk pages at multiple venues so having a centralised area for these discussions could be useful. Do editors think a taskforce page would be useful. @Tryptofish, Kingofaces43, Smartse, Lfstevens, Dialectric, and Tsavage: AIRcorn (talk) 07:29, 20 November 2017 (UTC)
- I think this is a good idea, and I've been intending to dome something with genome editing/genome engineering for a while. I think it would be sensible to keep the todo list well-defined and manageable so as not to lose steam. The GA blitz on the Gene article a few years back was very effective, so I could imagine a shortlist of articles being possible to overhaul (approx 1 article per 3-5 editors helping out). I can help out with images (and maybe some of the text). One additional feature that could be considered, invitation of experts on genetic engineering to help out, for example by emailing people who have authored good review articles (example), or written about GE in The Conversation (examples). It could be an ideal opportunity to attempt collaboration between experienced Wikipedians and specialist scientists. Also pinging: @Fixuture, Opabinia regalis, Boghog, Adrian J. Hunter, Jytdog, Splette, Samsara, Mark viking, Narayanese, Ettrig, Rjwilmsi, DennisPietras, Dcbennett2, and CatPath:. T.Shafee(Evo&Evo)talk 11:21, 20 November 2017 (UTC)
- Good idea. Lfstevens (talk) 15:43, 20 November 2017 (UTC)
- Not sure what to do here at this point other than say "hi" and maybe come back later. Samsara 18:11, 20 November 2017 (UTC)
- Thanks for the ping! I think it's potentially a very good idea and I wish it well. That said, my experience has been that "just do it" tends to work better than trying to herd
catseditors into anything organized. Genetics isn't my strong suit, and I'm frankly sick and tired of the whole GMO thing and eager to spend much less time on it, so I'm going to say no thank you to the invitation. But if there's a particular page where my help would be useful, please do not hesitate to ping me about it. --Tryptofish (talk) 20:17, 20 November 2017 (UTC)- I can't say I would be a lot of help as a primary contributor for the time being either as I'm on a bit of burnout in the subject too, and I usually only have heavy bursts of article editing on the rare occasions I have a good block of time to set aside (though I have a to-do list to tackle on the subject). That being said, I definitely am open to help out where I can for a review of content, etc. as an entomologist in crop protection. I keep up with crop genetics from way back in my grad school training and at scientific meetings still, so I'm happy to act as a subject-matter resource in that topic. If we get a list together of people in the task-force and what areas they would be good to ping on, I'm in for that capacity. Kingofaces43 (talk) 03:23, 21 November 2017 (UTC)
- Thanks for the ping! I think it's potentially a very good idea and I wish it well. That said, my experience has been that "just do it" tends to work better than trying to herd
Here is a possibly relevant collection of pageview stats on what I think are some of the most viewed genetic engineering articles. T.Shafee(Evo&Evo)talk 11:02, 22 November 2017 (UTC)
- Thank you everyone. There is enough interest here that I think it is worth starting this. Will see if and how it evolves. AIRcorn (talk) 08:59, 25 November 2017 (UTC)
Thanks for the consideration. I'm only on here rarely and randomly, lately. As for GE, it took just a refresher look at the language locked into a suite of GM articles by that ARBCOM, for me to want to have zero to do with the topic for the time being -- the consensus process seems too gameable and taxing to make contributing in any way satisfying (maybe later, whenever there are more broadly conclusive developments in the general subject area). Cheers! :) --Tsavage (talk) 13:08, 28 November 2017 (UTC)
Disambiguation links on pages tagged by this wikiproject
Wikipedia has many thousands of wikilinks which point to disambiguation pages. It would be useful to readers if these links directed them to the specific pages of interest, rather than making them search through a list. Members of WikiProject Disambiguation have been working on this and the total number is now below 20,000 for the first time. Some of these links require specialist knowledge of the topics concerned and therefore it would be great if you could help in your area of expertise.
A list of the relevant links on pages which fall within the remit of this wikiproject can be found at http://69.142.160.183/~dispenser/cgi-bin/topic_points.py?banner=WikiProject_Genetics
Please take a few minutes to help make these more useful to our readers.— Rod talk 15:47, 3 December 2017 (UTC)
- All
 Fixed! The one remaining dablink is intentional. Thanks Rodw! Adrian J. Hunter(talk•contribs) 01:42, 4 December 2017 (UTC)
Fixed! The one remaining dablink is intentional. Thanks Rodw! Adrian J. Hunter(talk•contribs) 01:42, 4 December 2017 (UTC)
- Thanks.— Rod talk 08:01, 4 December 2017 (UTC)
WikiCV for contributors
Hi all,
I'm a 4th year Computer Science undergraduate student based out of India. I've been selected as an intern for Wikimedia under Round-15 of Outreachy. I'll be building a web tool called WikiCV (under the mentorship of Gergő Tisza and Stephen LaPorte), somewhat similar to your LinkedIn, StackOverflow or Github profile. Before starting with the project, I wanted inputs from users who are my target audience, that is you all! So thought of asking it over the talk page. (My mentors pointed me to this page as here many ardent users are present.)
We came up with this project because we feel that Wikipedia needs a powerful force to draw new editors to the project and allow existing editors to spend more time on it without harming their career; unfortunately, due to the highly collaborative nature of Wikipedia, the value of one's participation is hard to measure for an outsider, which makes it very hard for contributors to take credit for value added to Wikipedia.
Hence, we want to create a contribution summarizing tool which (unlike the existing ones that focus on statistics and are hard to interpret for someone not familiar with Wikipedia editing) highlights contributions in an easy-to-understand manner.
I want your inputs on:
1. What all things would you like to see in your CV for Wikipedia contributions?
2. In what way should we present the data/ contribution summary so that it is understandable by a non-Wikipedia user?
3. What are the benefits/problems of the current tools that summarize the contribution of a user (like Xtools)?
4. The CV will definitely reflect your contribution, but would it be better if it shows your current standing with respect to other users? For example, reputation points in Stack Overflow reflect how good you are relatively. One idea that I thought was - Imagine a tool that tells someone is in the top 1% of editors. Would it be nice? If yes, what would you consider a good basis for that statement?
Also, I prepared a mockup for the CV to give a rough idea as to what we are thinking of. Please check it out as well.
Apart from this, I thought of presenting the contributions in a manner similar to Github. I've prepared a tool for that. Kindly have a look at that as well and give your reviews about it.
My work is largely dependent on your inputs, so please pour in your comments/views. Your help will be quite appreciated!
Anyone can reach out to me through mail(meghasharma4910@gmail.com) as well.
Eagerly waiting for your inputs :)
Meghasharma213 (talk) 15:56, 11 December 2017 (UTC)
- @Meghasharma213: This is a great idea. I'm a strong proponent of the idea of being able to raise the profile of Wikipedia as professional service to the community, and this sort of formal recognition framework is extremely useful.[1][2][3][4][5] I've a few thoughts that might help:
- Integrate and as much as possible with WikiData and draw ideas from existing tools
- Scholia - WikiData driven author analytics
- XTools and dewkin - Overall summary of edits from a very Wikipedian point of view
- AltMetric reports - An exiting measure of impact outside of narrow academic literature
- ORCID - Probably the best open standard for unique author IDs, (speak to Andy Mabbett)
- Whocolour - A great way of indicating an editors contributions to a key article
- Different Wikipedians contribute in different ways, consider multiple summary statistics, e.g.:
- Top 10 articles that the author has contributed to (for authors who contribute to big articles, but are never the dominant editor)
- Total number of articles where >50% of text is by the contributor (for authors that are the main contributors to a small number of articles)
- Total number of bytes / words / edits / different articles (for authors who contribute a little bit to a wide array of articles)
- Number of Good Articles and Featured articles
- Number of images, or a set of pinned images
- Not all edits are equal, different edits can signify very different habits
- Wordcloud or keywords of topics mentioned in abstracts of the top 20 articles they've edited?
- Main WikiProjects contributed to (main articles, article talk pages, WikiProject talk pages)
- Average bytes per edit? (put in percentile context)
- Average number of images per word? (put in percentile context)
- Some final thoughts:
- Some of these are computationally expensive (e.g. this tool takes ages) so consider ways of making it only need to update a cached version since the last time it was looked at.
- Avoid semi-3D bar graphs (flat is always better)
- Avoid stacked bar charts (overlayed line charts are usually better)[6]
- Make summary data downloadable
- Make clear when the page's data was last updated
- Allow users to choose which bits of data are shown
- For pinned articles, indicate percentage contribution by bytes
- For pinned articles, perhaps even implement something like WhoColor
- An area to list all other online presences (again Wikidata integrated) e.g. LinkedIn, ResearchGate, G-scholar, Mendelay, Academia.com, Faculty page, Homepage, Userpage, etc
- Integrate and as much as possible with WikiData and draw ideas from existing tools
- Best of luck. I hope any of that is helpful, and I look forward to following the project as it develops! T.Shafee(Evo&Evo)talk 09:38, 12 December 2017 (UTC)
- @Meghasharma213: This is a great idea. I'm a strong proponent of the idea of being able to raise the profile of Wikipedia as professional service to the community, and this sort of formal recognition framework is extremely useful.[1][2][3][4][5] I've a few thoughts that might help:
References
- ^ Allen, Liz; Scott, Jo; Brand, Amy; Hlava, Marjorie; Altman, Micah (2014-04-17). "Publishing: Credit where credit is due". Nature. 508 (7496): 312–313. Bibcode:2014Natur.508..312A. doi:10.1038/508312a. PMID 24745070.
- ^ Narayan, Sneha; Orlowitz, Jake; Morgan, Jonathan T.; Shaw, Aaron (2015). "Effects of a Wikipedia Orientation Game on New User Edits". Proceedings of the 18th ACM Conference Companion on Computer Supported Cooperative Work & Social Computing. CSCW'15 Companion. New York, NY, USA: ACM. pp. 263–266. doi:10.1145/2685553.2699022. ISBN 9781450329460. S2CID 18534458.
- ^ Sugimoto, Cassidy R.; Work, Sam; Larivière, Vincent; Haustein, Stefanie (2017-09-01). "Scholarly use of social media and altmetrics: A review of the literature". Journal of the Association for Information Science and Technology. 68 (9): 2037–2062. arXiv:1608.08112. doi:10.1002/asi.23833. ISSN 2330-1643. S2CID 1572378.
- ^ Arazy, Ofer; Stroulia, Eleni; Ruecker, Stan; Arias, Cristina; Fiorentino, Carlos; Ganev, Veselin; Yau, Timothy (2010-06-01). "Recognizing contributions in wikis: Authorship categories, algorithms, and visualizations". Journal of the American Society for Information Science and Technology. 61 (6): 1166–1179. doi:10.1002/asi.21326. ISSN 1532-2890.
- ^ Shafee, Thomas; Mietchen, Daniel; Su, Andrew I. (2017-08-11). "Academics can help shape Wikipedia". Science. 357 (6351): 557–558. Bibcode:2017Sci...357..557S. doi:10.1126/science.aao0462. ISSN 0036-8075. PMID 28798122. S2CID 19075849.
- ^ Rougier, Nicolas P.; Droettboom, Michael; Bourne, Philip E. (2014-09-11). "Ten Simple Rules for Better Figures". PLOS Computational Biology. 10 (9): e1003833. Bibcode:2014PLSCB..10E3833R. doi:10.1371/journal.pcbi.1003833. ISSN 1553-7358. PMID 25210732.
Thanks a lot T.Shafee(Evo&Evo) for your comments. Quite insightful, I must say! Will keep you updated about the project. Meghasharma213 (talk) 14:32, 13 December 2017 (UTC)
- @Meghasharma213: Thanks, I'm particularly interested, because I think that it could be applied to the authors of WikiJournal (e.g. in Wiki.J.Med.). T.Shafee(Evo&Evo)talk 22:36, 14 December 2017 (UTC)
Human Disease Modifier Gene
Could someone have a look at Draft:Human Disease Modifier Gene and see if its sourcing is reliable enough for it to be moved to mainspace? Many thanks! – Uanfala (talk) 17:16, 7 January 2018 (UTC)
- That draft requires more time than I am willing to donate, but I think the subject is notorious for WP. In particular, the lede section has to be remade and soften the scientific terms to reach high-school-level readers. BatteryIncluded (talk) 18:00, 7 January 2018 (UTC)
- I made an attempt to make the lead more accessible. It needs more work, but it may be adequate to move to main space. Boghog (talk) 07:45, 8 January 2018 (UTC)
- Good! I'm moving to mainspace then. Thank you for improving it! – Uanfala (talk) 19:58, 9 January 2018 (UTC)
- I made an attempt to make the lead more accessible. It needs more work, but it may be adequate to move to main space. Boghog (talk) 07:45, 8 January 2018 (UTC)
Links to DAB pages
There may be a hundred or so genetics-related articles which contain bad links to DAB pages. User:DPL bot logs them; I can find them; you need do no searching; I think I know enough about genetics to know when not to guess; any help would be appreciated (most of all, by our readers).
In this list, search for "disam" in the text as displayed. If you solve a problem, remove ((dn)) from the article and add ((done)) here. It's possible that some of them may need appropriate redlinking (we all know that there are gaps in Wiki).
- BC200 lncRNA
 Done
Done - Alternative splicing
 Done by a previous editor
Done by a previous editor - The Monarch Initiative
- DNA (cytosine-5)-methyltransferase 3A
 Done by a previous editor
Done by a previous editor - SH3KBP1
 Done
Done - Razi Vaccine and Serum Research Institute
 Done
Done - IgSF CAM
 Done
Done
That's just an initial batch. Thanks in advance for any help. Narky Blert (talk) 22:49, 24 January 2018 (UTC)
- Not sure what to choose for The Monarch Initiative; it seems that all of the mechanisms of 'genetic variant' would apply. Leschnei (talk) 14:43, 3 February 2018 (UTC)
There appears to be a close relationship between these two articles which probably requires a summary section and wikilinks at both ends. The maternal effect is complex and debatable, and telegony antique, so the situation demands some expertise. I'd be glad of any assistance. Chiswick Chap (talk) 10:19, 22 January 2018 (UTC)
- Chiswick Chap: as I understand things, the effects are quite different. The first is supported by evidence, the second is not. Maproom (talk) 14:21, 5 February 2018 (UTC)
Facto Post – Issue 10 – 12 March 2018
Facto Post – Issue 10 – 12 March 2018
 Milestone for mix'n'matchAround the time in February when Wikidata clicked past item Q50000000, another milestone was reached: the mix'n'match tool uploaded its 1000th dataset. Concisely defined by its author, Magnus Manske, it works "to match entries in external catalogs to Wikidata". The total number of entries is now well into eight figures, and more are constantly being added: a couple of new catalogs each day is normal. Since the end of 2013, mix'n'match has gradually come to play a significant part in adding statements to Wikidata. Particularly in areas with the flavour of digital humanities, but datasets can of course be about practically anything. There is a catalog on skyscrapers, and two on spiders. These days mix'n'match can be used in numerous modes, from the relaxed gamified click through a catalog looking for matches, with prompts, to the fantastically useful and often demanding search across all catalogs. I'll type that again: you can search 1000+ datasets from the simple box at the top right. The drop-down menu top left offers "creation candidates", Magnus's personal favourite. m:Mix'n'match/Manual for more. For the Wikidatan, a key point is that these matches, however carried out, add statements to Wikidata if, and naturally only if, there is a Wikidata property associated with the catalog. For everyone, however, the hands-on experience of deciding of what is a good match is an education, in a scholarly area, biographical catalogs being particularly fraught. Underpinning recent rapid progress is an open infrastructure for scraping and uploading. Congratulations to Magnus, our data Stakhanovite! Links
Editor Charles Matthews, for ContentMine. Please leave feedback for him. Back numbers are here. Reminder: WikiFactMine pages on Wikidata are at WD:WFM. If you wish to receive no further issues of Facto Post, please remove your name from our mailing list. Alternatively, to opt out of all massmessage mailings, you may add Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery (talk) 12:26, 12 March 2018 (UTC)
Behavioural genetics
Behavioural genetics, an article that you or your project may be interested in, has been nominated for a community good article reassessment. If you are interested in the discussion, please participate by adding your comments to the reassessment page. If concerns are not addressed during the review period, the good article status may be removed from the article. AIRcorn (talk) 09:48, 20 March 2018 (UTC)
Merger proposal related to Lábrea fever and Hepatitis D
A discussion is taking place that may be of interest to some members of this project at Talk:Hepatitis_D#Merger_proposal. — soupvector (talk) 13:42, 2 April 2018 (UTC)
I work at a genomics company called Clear Labs. I have disclosed my conflict of interest on the Talk page of the Clear Labs article and offered a proposed replacement of the article that is much shorter, less promotional, less out-of-date, better-sourced, etc. here. It’s been a couple weeks with no response on the article’s Talk page, so I wanted to see if anyone here had a minute to review my proposed draft and let me know if I was doing anything inappropriate COI-wise. Mark Bajus at Clear Labs (talk) 22:33, 5 April 2018 (UTC)
![]() Done Natureium (talk) 17:27, 6 April 2018 (UTC)
Done Natureium (talk) 17:27, 6 April 2018 (UTC)
Any interest in rescuing this abandoned draft? Espresso Addict (talk) 01:25, 9 April 2018 (UTC)
- Cleaned it up and moved to main space. As always, could use some more editing. Boghog (talk) 04:35, 9 April 2018 (UTC)
- Thank you! Always good to see content assimilated. Espresso Addict (talk) 05:22, 9 April 2018 (UTC)
- Well done. I'll also contact a few academics to see if I can convince any to write an article on Paramecium tetraurelia. T.Shafee(Evo&Evo)talk 07:02, 9 April 2018 (UTC)
- Thank you! Always good to see content assimilated. Espresso Addict (talk) 05:22, 9 April 2018 (UTC)
Facto Post – Issue 11 – 9 April 2018
Facto Post – Issue 11 – 9 April 2018
 The 100 Skins of the OnionOpen Citations Month, with its eminently guessable hashtag, is upon us. We should be utterly grateful that in the past 12 months, so much data on which papers cite which other papers has been made open, and that Wikidata is playing its part in hosting it as "cites" statements. At the time of writing, there are 15.3M Wikidata items that can do that. Pulling back to look at open access papers in the large, though, there is is less reason for celebration. Access in theory does not yet equate to practical access. A recent LSE IMPACT blogpost puts that issue down to "heterogeneity". A useful euphemism to save us from thinking that the whole concept doesn't fall into the realm of the oxymoron. Some home truths: aggregation is not content management, if it falls short on reusability. The PDF file format is wedded to how humans read documents, not how machines ingest them. The salami-slicer is our friend in the current downloading of open access papers, but for a better metaphor, think about skinning an onion, laboriously, 100 times with diminishing returns. There are of the order of 100 major publisher sites hosting open access papers, and the predominant offer there is still a PDF.  From the discoverability angle, Wikidata's bibliographic resources combined with the SPARQL query are superior in principle, by far, to existing keyword searches run over papers. Open access content should be managed into consistent HTML, something that is currently strenuous. The good news, such as it is, would be that much of it is already in XML. The organisational problem of removing further skins from the onion, with sensible prioritisation, is certainly not insuperable. The CORE group (the bloggers in the LSE posting) has some answers, but actually not all that is needed for the text and data mining purposes they highlight. The long tail, or in other words the onion heart when it has become fiddly beyond patience to skin, does call for a pis aller. But the real knack is to do more between the XML and the heart. Links
Editor Charles Matthews, for ContentMine. Please leave feedback for him. Back numbers are here. Reminder: WikiFactMine pages on Wikidata are at WD:WFM. If you wish to receive no further issues of Facto Post, please remove your name from our mailing list. Alternatively, to opt out of all massmessage mailings, you may add Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery (talk) 16:25, 9 April 2018 (UTC)
Automatic archiving of this page again?
Hi folks, it seems like this page used to be automatically archived by lowercase sigmabot III, but that stopped about a year and a half ago when we switched to the sleeker talk page heading at the top. I don't quite understand how the new talk navbox works, but is there a way to set up automatic talk page archiving again (alternatively, is that something folks want?)? Thanks Ajpolino (talk) 18:07, 9 April 2018 (UTC)
- I think that the new header should be compatible with lowercase sigmabot III (since the bot creates the archive subpages, and the header just needs to link to the created subpages). I've therefore re-added the ((User:MiszaBot/config)) configuration template. T.Shafee(Evo&Evo)talk 23:39, 9 April 2018 (UTC)
Proposal: Merge 'Exome' and 'Coding Region' Harveyjamesm (talk) 21:51, 10 May 2018 (UTC)
Exome and Coding region are nearly synonymous. You could argue "coding region" might refer specifically to a single gene whereas "exome" is global, but wouldn't it be more logical to put them both together? I'd personally argue Exome is a more up to date descriptor, plus the exome consists of many coding regions, and is in fact the coding region of the entire genome.Harveyjamesm (talk) 21:51, 10 May 2018 (UTC)
- Thanks for the suggestion, but they are not the same concept. An exome is the total set of exons in an organism. An exon is a coding region plus any flanking UTRs. Hence coding regions form an important part of the exome, but are not synonymous with it. --Mark viking (talk) 22:30, 10 May 2018 (UTC)
- thanks for clarifying that Harveyjamesm (talk) 08:34, 11 May 2018 (UTC)
- I think that the topics are sufficiently different to have separate pages. Protein coding region is most commonly used to refer to regions of individual genes rather than those regions across the whole genome. T.Shafee(Evo&Evo)talk 10:04, 11 May 2018 (UTC)
Facto Post – Issue 12 – 28 May 2018
Facto Post – Issue 12 – 28 May 2018
 ScienceSource fundedThe Wikimedia Foundation announced full funding of the ScienceSource grant proposal from ContentMine on May 18. See the ScienceSource Twitter announcement and 60 second video.
The proposal includes downloading 30,000 open access papers, aiming (roughly speaking) to create a baseline for medical referencing on Wikipedia. It leaves open the question of how these are to be chosen. The basic criteria of WP:MEDRS include a concentration on secondary literature. Attention has to be given to the long tail of diseases that receive less current research. The MEDRS guideline supposes that edge cases will have to be handled, and the premature exclusion of publications that would be in those marginal positions would reduce the value of the collection. Prophylaxis misses the point that gate-keeping will be done by an algorithm. Two well-known but rather different areas where such considerations apply are tropical diseases and alternative medicine. There are also a number of potential downloading troubles, and these were mentioned in Issue 11. There is likely to be a gap, even with the guideline, between conditions taken to be necessary but not sufficient, and conditions sufficient but not necessary, for candidate papers to be included. With around 10,000 recognised medical conditions in standard lists, being comprehensive is demanding. With all of these aspects of the task, ScienceSource will seek community help. Links
Editor Charles Matthews, for ContentMine. Please leave feedback for him. Back numbers are here. Reminder: WikiFactMine pages on Wikidata are at WD:WFM. ScienceSource pages will be announced there, and in this mass message. If you wish to receive no further issues of Facto Post, please remove your name from our mailing list. Alternatively, to opt out of all massmessage mailings, you may add Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery (talk) 10:16, 28 May 2018 (UTC)
Facto Post – Issue 13 – 29 May 2018
Facto Post – Issue 13 – 29 May 2018
 The Editor is Charles Matthews, for ContentMine. Please leave feedback for him, on his User talk page.
To subscribe to Facto Post go to Wikipedia:Facto Post mailing list. For the ways to unsubscribe, see the footer.
Facto Post enters its second year, with a Cambridge Blue (OK, Aquamarine) background, a new logo, but no Cambridge blues. On-topic for the ScienceSource project is a project page here. It contains some case studies on how the WP:MEDRS guideline, for the referencing of articles at all related to human health, is applied in typical discussions. Close to home also, a template, called ((medrs)) for short, is used to express dissatisfaction with particular references. Technology can help with patrolling, and this Petscan query finds over 450 articles where there is at least one use of the template. Of course the template is merely suggesting there is a possible issue with the reliability of a reference. Deciding the truth of the allegation is another matter. This maintenance issue is one example of where ScienceSource aims to help. Where the reference is to a scientific paper, its type of algorithm could give a pass/fail opinion on such references. It could assist patrollers of medical articles, therefore, with the templated references and more generally. There may be more to proper referencing than that, indeed: context, quite what the statement supported by the reference expresses, prominence and weight. For that kind of consideration, case studies can help. But an algorithm might help to clear the backlog. 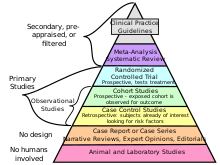
If you wish to receive no further issues of Facto Post, please remove your name from our mailing list. Alternatively, to opt out of all massmessage mailings, you may add Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery (talk) 18:19, 29 June 2018 (UTC)
Facto Post – Issue 14 – 21 July 2018
Facto Post – Issue 14 – 21 July 2018
 The Editor is Charles Matthews, for ContentMine. Please leave feedback for him, on his User talk page.
To subscribe to Facto Post go to Wikipedia:Facto Post mailing list. For the ways to unsubscribe, see the footer.
Officially it is "bridging the gaps in knowledge", with Wikimania 2018 in Cape Town paying tribute to the southern African concept of ubuntu to implement it. Besides face-to-face interactions, Wikimedians do need their power sources.  Facto Post interviewed Jdforrester, who has attended every Wikimania, and now works as Senior Product Manager for the Wikimedia Foundation. His take on tackling the gaps in the Wikimedia movement is that "if we were an army, we could march in a column and close up all the gaps". In his view though, that is a faulty metaphor, and it leads to a completely false misunderstanding of the movement, its diversity and different aspirations, and the nature of the work as "fighting" to be done in the open sector. There are many fronts, and as an eventualist he feels the gaps experienced both by editors and by users of Wikimedia content are inevitable. He would like to see a greater emphasis on reuse of content, not simply its volume. If that may not sound like radicalism, the Decolonizing the Internet conference here organized jointly with Whose Knowledge? can redress the picture. It comes with the claim to be "the first ever conference about centering marginalized knowledge online". 
If you wish to receive no further issues of Facto Post, please remove your name from our mailing list. Alternatively, to opt out of all massmessage mailings, you may add Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery (talk) 06:10, 21 July 2018 (UTC)
Facto Post – Issue 15 – 21 August 2018
Facto Post – Issue 15 – 21 August 2018
 The Editor is Charles Matthews, for ContentMine. Please leave feedback for him, on his User talk page.
To subscribe to Facto Post go to Wikipedia:Facto Post mailing list. For the ways to unsubscribe, see the footer.
 To grasp the nettle, there are rare diseases, there are tropical diseases and then there are "neglected diseases". Evidently a rare enough disease is likely to be neglected, but neglected disease these days means a disease not rare, but tropical, and most often infectious or parasitic. Rare diseases as a group are dominated, in contrast, by genetic diseases. A major aspect of neglect is found in tracking drug discovery. Orphan drugs are those developed to treat rare diseases (rare enough not to have market-driven research), but there is some overlap in practice with the WHO's neglected diseases, where snakebite, a "neglected public health issue", is on the list. From an encyclopedic point of view, lack of research also may mean lack of high-quality references: the core medical literature differs from primary research, since it operates by aggregating trials. This bibliographic deficit clearly hinders Wikipedia's mission. The ScienceSource project is currently addressing this issue, on Wikidata. Its Wikidata focus list at WD:SSFL is trying to ensure that neglect does not turn into bias in its selection of science papers.
If you wish to receive no further issues of Facto Post, please remove your name from our mailing list. Alternatively, to opt out of all massmessage mailings, you may add Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery (talk) 13:23, 21 August 2018 (UTC)
Request for comments
This is a request for comments regarding the relevance of Denny (hybrid hominin) as it relates to human interspecies breeding. The link to the discussion is at: Talk:Human evolution#A curious discovery, but not really on-topic. Cheers, Rowan Forest (talk) 16:37, 26 August 2018 (UTC)
Sex determination material at the Sexual differentiation in humans article
Opinions are needed on the following: Talk:Sexual differentiation in humans#"A baby’s sex is determined at the time of conception." citing 17th edition of Harrison's. The discussion concerns how to add material such as females typically having two X chromosomes (XX), and other material. A permalink for it is here. Flyer22 Reborn (talk) 21:24, 31 August 2018 (UTC)
RfC being planned
Please see WP:Reliable_sources/Noticeboard#Primary_genetics_studies. Jytdog (talk) 23:40, 6 September 2018 (UTC)
This is an interesting article with reliable sources SebaSebaBukh (talk) 21:24, 9 September 2018 (UTC)
Facto Post – Issue 16 – 30 September 2018
Facto Post – Issue 16 – 30 September 2018
 The Editor is Charles Matthews, for ContentMine. Please leave feedback for him, on his User talk page.
To subscribe to Facto Post go to Wikipedia:Facto Post mailing list. For the ways to unsubscribe, see the footer.
 In an ideal world ... no, bear with your editor for just a minute ... there would be a format for scientific publishing online that was as much a standard as SI units are for the content. Likewise cataloguing publications would not be onerous, because part of the process would be to generate uniform metadata. Without claiming it could be the mythical free lunch, it might be reasonably be argued that sandwiches can be packaged much alike and have barcodes, whatever the fillings. The best on offer, to stretch the metaphor, is the meal kit option, in the form of XML. Where scientific papers are delivered as XML downloads, you get all the ingredients ready to cook. But have to prepare the actual meal of slow food yourself. See Scholarly HTML for a recent pass at heading off XML with HTML, in other words in the native language of the Web. The argument from real life is a traditional mixture of frictional forces, vested interests, and the classic irony of the principle of unripe time. On the other hand, discoverability actually diminishes with the prolific progress of science publishing. No, it really doesn't scale. Wikimedia as movement can do something in such cases. We know from open access, we grok the Web, we have our own horse in the HTML race, we have Wikidata and WikiJournal, and we have the chops to act. 
If you wish to receive no further issues of Facto Post, please remove your name from our mailing list. Alternatively, to opt out of all massmessage mailings, you may add Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery (talk) 17:57, 30 September 2018 (UTC)
Comment on draft
Your comments on Draft:Haplogroup E-M81 are welcomed. Please use either Yet Another Articles for Creation Helper Script by enabling Preferences → Gadgets → Editing → ![]() Yet Another AFC Helper Script, or use
Yet Another AFC Helper Script, or use ((afc comment|Your comment here. ~~~~)) directly in the draft. Thank you. Sam Sailor 19:06, 28 October 2018 (UTC)
Facto Post – Issue 17 – 29 October 2018
Facto Post – Issue 17 – 29 October 2018
 The Editor is Charles Matthews, for ContentMine. Please leave feedback for him, on his User talk page.
To subscribe to Facto Post go to Wikipedia:Facto Post mailing list. For the ways to unsubscribe, see the footer.
Around 2.7 million Wikidata items have an illustrative image. These files, you might say, are Wikimedia's stock images, and if the number is large, it is still only 5% or so of items that have one. All such images are taken from Wikimedia Commons, which has 50 million media files. One key issue is how to expand the stock. Indeed, there is a tool. WD-FIST exploits the fact that each Wikipedia is differently illustrated, mostly with images from Commons but also with fair use images. An item that has sitelinks but no illustrative image can be tested to see if the linked wikis have a suitable one. This works well for a volunteer who wants to add images at a reasonable scale, and a small amount of SPARQL knowledge goes a long way in producing checklists.  It should be noted, though, that there are currently 53 Wikidata properties that link to Commons, of which P18 for the basic image is just one. WD-FIST prompts the user to add signatures, plaques, pictures of graves and so on. There are a couple of hundred monograms, mostly of historical figures, and this query allows you to view all of them. commons:Category:Monograms and its subcategories provide rich scope for adding more. And so it is generally. The list of properties linking to Commons does contain a few that concern video and audio files, and rather more for maps. But it contains gems such as P3451 for "nighttime view". Over 1000 of those on Wikidata, but as for so much else, there could be yet more. Go on. Today is Wikidata's birthday. An illustrative image is always an acceptable gift, so why not add one? You can follow these easy steps: (i) log in at https://tools.wmflabs.org/widar/, (ii) paste the Petscan ID 6263583 into https://tools.wmflabs.org/fist/wdfist/ and click run, and (iii) just add cake. 
If you wish to receive no further issues of Facto Post, please remove your name from our mailing list. Alternatively, to opt out of all massmessage mailings, you may add Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery (talk) 15:01, 29 October 2018 (UTC)
Featured quality source review RFC
Editors in this WikiProject may be interested in the featured quality source review RFC that has been ongoing. It would change the featured article candidate process (FAC) so that source reviews would need to occur prior to any other reviews for FAC. Your comments are appreciated. --IznoRepeat (talk) 21:50, 11 November 2018 (UTC)
Request for a second opinion on Cytohet
Hi WP Genetics members, is Cytohet notable enough for its own article in your opinion? Genetics is not my strong point, but it is currently tagged ((notability)). Thanks! --TheSandDoctor Talk 19:25, 12 November 2018 (UTC)
- I think it is probably notable, but the proper home for the article is likely "heteroplasmon", rather than cytohet. Canada Hky (talk) 13:46, 13 November 2018 (UTC)
Topic Page on Selfish genetic elements
PLOS Genetics has now joined PLOS Computational Biology in its Topic Pages initiative. As part of this, an article was drafted, peer reviewed and published in PLOS Genetics and has now been copied over to the Selfish genetic element wikipedia page. Comments and suggestions welcome! T.Shafee(Evo&Evo)talk 00:41, 17 November 2018 (UTC)
Facto Post – Issue 18 – 30 November 2018
Facto Post – Issue 18 – 30 November 2018
 The Editor is Charles Matthews, for ContentMine. Please leave feedback for him, on his User talk page.
To subscribe to Facto Post go to Wikipedia:Facto Post mailing list. For the ways to unsubscribe, see the footer.
GLAM ♥ data — what is a gallery, library, archive or museum without a catalogue? It follows that Wikidata must love librarians. Bibliography supports students and researchers in any topic, but open and machine-readable bibliographic data even more so, outside the silo. Cue the WikiCite initiative, which was meeting in conference this week, in the Bay Area of California.  In fact there is a broad scope: "Open Knowledge Maps via SPARQL" and the "Sum of All Welsh Literature", identification of research outputs, Library.Link Network and Bibframe 2.0, OSCAR and LUCINDA (who they?), OCLC and Scholia, all these co-exist on the agenda. Certainly more library science is coming Wikidata's way. That poses the question about the other direction: is more Wikimedia technology advancing on libraries? Good point. Wikimedians generally are not aware of the tech background that can be assumed, unless they are close to current training for librarians. A baseline definition is useful here: "bash, git and OpenRefine". Compare and contrast with pywikibot, GitHub and mix'n'match. Translation: scripting for automation, version control, data set matching and wrangling in the large, are on the agenda also for contemporary library work. Certainly there is some possible common ground here. Time to understand rather more about the motivations that operate in the library sector.
Account creation is now open on the ScienceSource wiki, where you can see SPARQL visualisations of text mining.
If you wish to receive no further issues of Facto Post, please remove your name from our mailing list. Alternatively, to opt out of all massmessage mailings, you may add Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery (talk) 11:20, 30 November 2018 (UTC)
Request for assessment on MiR-324-5p -Dec 11 2018
The page https://en.wikipedia.org/wiki/MiR-324-5p was developed as part of course on science communication at the University of Cincinnati. It has not been rated yet for importance or quality. It would be great if someone could help with that. thanks! TMorata (talk) 12:27, 11 December 2018 (UTC)TMorata
Topic Page on Micropeptides
PLOS Genetics just published its second Topic Pages article. As with the previous one on selfish genetic elements, it was drafted, peer reviewed and published in PLOS Genetics and has now been copied over to the Micropeptide wikipedia page. Comments, suggestions, edits and updates welcome! T.Shafee(Evo&Evo)talk 12:45, 14 December 2018 (UTC)
Facto Post – Issue 19 – 27 December 2018
Facto Post – Issue 19 – 27 December 2018
 The Editor is Charles Matthews, for ContentMine. Please leave feedback for him, on his User talk page.
To subscribe to Facto Post go to Wikipedia:Facto Post mailing list. For the ways to unsubscribe, see the footer.
Zotero is free software for reference management by the Center for History and New Media: see Wikipedia:Citing sources with Zotero. It is also an active user community, and has broad-based language support. Besides the handiness of Zotero's warehousing of personal citation collections, the Zotero translator underlies the citoid service, at work behind the VisualEditor. Metadata from Wikidata can be imported into Zotero; and in the other direction the zotkat tool from the University of Mannheim allows Zotero bibliographies to be exported to Wikidata, by item creation. With an extra feature to add statements, that route could lead to much development of the focus list (P5008) tagging on Wikidata, by WikiProjects. There is also a large-scale encyclopedic dimension here. The construction of Zotero translators is one facet of Web scraping that has a strong community and open source basis. In that it resembles the less formal mix'n'match import community, and growing networks around other approaches that can integrate datasets into Wikidata, such as the use of OpenRefine. Looking ahead, the thirtieth birthday of the World Wide Web falls in 2019, and yet the ambition to make webpages routinely readable by machines can still seem an ever-retreating mirage. Wikidata should not only be helping Wikimedia integrate its projects, an ongoing process represented by Structured Data on Commons and lexemes. It should also be acting as a catalyst to bring scraping in from the cold, with institutional strengths as well as resourceful code.
Diversitech, the latest ContentMine grant application to the Wikimedia Foundation, is in its community review stage until January 2.
If you wish to receive no further issues of Facto Post, please remove your name from our mailing list. Alternatively, to opt out of all massmessage mailings, you may add Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery (talk) 19:08, 27 December 2018 (UTC)
Discussion at RSN on the use of eupedia.com for genetics
It seems to be a one man band, started off as a tourist site and the site owner became interested in genetics. Doug Weller talk 20:31, 10 January 2019 (UTC)
- It looks like the link for that discussion is at Wikipedia:Reliable_sources/Noticeboard#Eupedia.com. --
((u|Mark viking)) {Talk}20:56, 10 January 2019 (UTC)
Facto Post – Issue 20 – 31 January 2019
Facto Post – Issue 20 – 31 January 2019
 The Editor is Charles Matthews, for ContentMine. Please leave feedback for him, on his User talk page.
To subscribe to Facto Post go to Wikipedia:Facto Post mailing list. For the ways to unsubscribe, see the footer.
Recently Jimmy Wales has made the point that computer home assistants take much of their data from Wikipedia, one way or another. So as well as getting Spotify to play Frosty the Snowman for you, they may be able to answer the question "is the Pope Catholic?" Possibly by asking for disambiguation (Coptic?). Headlines about data breaches are now familiar, but the unannounced circulation of information raises other issues. One of those is Gresham's law stated as "bad data drives out good". Wikipedia and now Wikidata have been criticised on related grounds: what if their content, unattributed, is taken to have a higher standing than Wikimedians themselves would grant it? See Wikiquote on a misattribution to Bismarck for the usual quip about "law and sausages", and why one shouldn't watch them in the making. Wikipedia has now turned 18, so should act like as adult, as well as being treated like one. The Web itself turns 30 some time between March and November this year, per Tim Berners-Lee. If the Knowledge Graph by Google exemplifies Heraclitean Web technology gaining authority, contra GIGO, Wikimedians still have a role in its critique. But not just with the teenage skill of detecting phoniness. There is more to beating Gresham than exposing the factoid and urban myth, where WP:V does do a great job. Placeholders must be detected, and working with Wikidata is a good way to understand how having one statement as data can blind us to replacing it by a more accurate one. An example that is important to open access is that, firstly, the term itself needs considerable unpacking, because just being able to read material online is a poor relation of "open"; and secondly, trying to get Creative Commons license information into Wikidata shows up issues with classes of license (such as CC-BY) standing for the actual license in major repositories. Detailed investigation shows that "everything flows" exacerbates the issue. But Wikidata can solve it.
If you wish to receive no further issues of Facto Post, please remove your name from our mailing list. Alternatively, to opt out of all massmessage mailings, you may add Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery (talk) 10:53, 31 January 2019 (UTC)
Facto Post – Issue 21 – 28 February 2019
Facto Post – Issue 21 – 28 February 2019
 The Editor is Charles Matthews, for ContentMine. Please leave feedback for him, on his User talk page.
To subscribe to Facto Post go to Wikipedia:Facto Post mailing list. For the ways to unsubscribe, see the footer.
Systematic reviews are basic building blocks of evidence-based medicine, surveys of existing literature devoted typically to a definite question that aim to bring out scientific conclusions. They are principled in a way Wikipedians can appreciate, taking a critical view of their sources.  Ben Goldacre in 2014 wrote (link below) "[...] : the "information architecture" of evidence based medicine (if you can tolerate such a phrase) is a chaotic, ad hoc, poorly connected ecosystem of legacy projects. In some respects the whole show is still run on paper, like it's the 19th century." Is there a Wikidatan in the house? Wouldn't some machine-readable content that is structured data help? Most likely it would, but the arcana of systematic reviews and how they add value would still need formal handling. The PRISMA standard dates from 2009, with an update started in 2018. The concerns there include the corpus of papers used: how selected and filtered? Now that Wikidata has a 20.9 million item bibliography, one can at least pose questions. Each systematic review is a tagging opportunity for a bibliography. Could that tagging be reproduced by a query, in principle? Can it even be second-guessed by a query (i.e. simulated by a protocol which translates into SPARQL)? Homing in on the arcana, do the inclusion and filtering criteria translate into metadata? At some level they must, but are these metadata explicitly expressed in the articles themselves? The answer to that is surely "no" at this point, but can TDM find them? Again "no", right now. Automatic identification doesn't just happen. Actually these questions lack originality. It should be noted though that WP:MEDRS, the reliable sources guideline used here for health information, hinges on the assumption that the usefully systematic reviews of biomedical literature can be recognised. Its nutshell summary, normally the part of a guideline with the highest density of common sense, allows literature reviews in general validity, but WP:MEDASSESS qualifies that indication heavily. Process wonkery about systematic reviews definitely has merit.
If you wish to receive no further issues of Facto Post, please remove your name from our mailing list. Alternatively, to opt out of all massmessage mailings, you may add Category:Wikipedians who opt out of message delivery to your user talk page.
Newsletter delivered by MediaWiki message delivery |
MediaWiki message delivery (talk) 10:02, 28 February 2019 (UTC)
