Mitogen-activated protein kinase kinase kinase 7 (MAP3K7), also known as TAK1, is an enzyme that in humans is encoded by the MAP3K7 gene.[5]
Structure
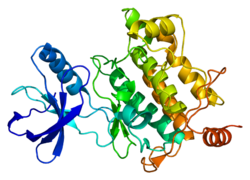
[edit]TAK1 is an evolutionarily conserved kinase in the MAP3 K family and clusters with the tyrosine-like and sterile kinase families. The protein structure of TAK1 contains an N (residues 1–104)- and C (residues 111–303)-terminus connected through the hinge region (Met 104-Ser 111). The ATP binding pocket is located in the hinge region of the kinase. Additionally, TAK1 has a catalytic lysine (Lys63) in the active site. Crystal structure of TAK1-ATP have shown that ATP forms two hydrogen bonds with residues Ala 107 and Glu 105. Further hydrogen bonding is observed to Asp 175, which is the leading residue of the DFG motif. This residue is thought to interact with Lys 63 through polar interactions and is catalytically important for phosphate transfer to substrate molecules. Critical for the TAK1-TAB1 complex is a helical loop around Phe 484, which provides extensive surface contact between the two proteins.
Signaling
[edit]The protein encoded by this gene is a member of the serine/threonine protein kinase family. This kinase mediates signal transduction induced by TGF beta and morphogenetic protein (BMP), and controls a variety of cell functions including transcription regulation and apoptosis. TAK1 is a central regulator of cell death and is activated through a diverse set of intra- and extracellular stimuli. TAK1 regulates cell survival not solely through NF-κB but also through NF-κB-independent pathways such as oxidative stress and receptor-interacting protein kinase 1 (RIPK1) kinase activity-dependent pathway.[6] In response to IL-1, this protein forms a kinase complex including TRAF6, MAP3K7P1/TAB1 and MAP3K7P2/TAB2; this complex is required for the activation of nuclear factor kappa B. This kinase can also activate MAPK8/JNK, MAP2K4/MKK4, and thus plays a role in the cell response to environmental stresses. Four alternatively spliced transcript variants encoding distinct isoforms have been reported.[7]]
In addition to IL-1 agonist activation, TAK1has been shown to be activated following TNF, TGFB, and LPS stimulation leading to the activation of pro-inflammatory pathways. Following TNF stimulation, TAK1 forms a ternary structure with TAB1 and TAB2/3 to form a fully activated TAK1 kinase. Following activation, TAK1 phosphorylates downstream effectors such as NFKB, p38 and cJUN leading the up-regulation of pro inflammatory pro survival genes.
Role in autoimmune diseases
[edit]This kinase has also been shown to regulate downstream cytokine expression such as TNF. Due to its regulation of TNF, TAK1 has become a novel target for the treatment of TNF mediated diseases such as auto immune disease ( Rheumatoid Arthritis, lupus, IBD) but also other cytokine mediated disorders such as chronic pain and cancer.[8][9] With the advent of novel selective TAK1 inhibitors, groups have explored the therapeutic potential of TAK1 targeted therapies. One group has shown that the selective TAK1 inhibitor, Takinib developed at Duke University attenuated rheumatoid arthritis like pathology in the CIA mouse model of human inflammatory arthritis.[10] Furthermore, pharmacological inhibition of TAK1 has shown to reduce inflammatory cytokines in particular TNF.[11]
Humans mutations
[edit]A rare mutation in TAK1 in humans has been reported. The mutation leads to gain of function, and hyper activation of TAK1 signaling pathways. Patients with gain of function mutations often present with craniofacial abnormalities.
Interactions
[edit]MAP3K7 has been shown to interact with:
References
[edit]- ^ a b c GRCh38: Ensembl release 89: ENSG00000135341 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000028284 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ Kondo M, Osada H, Uchida K, Yanagisawa K, Masuda A, Takagi K, et al. (February 1998). "Molecular cloning of human TAK1 and its mutational analysis in human lung cancer". International Journal of Cancer. 75 (4): 559–63. doi:10.1002/(SICI)1097-0215(19980209)75:4<559::AID-IJC11>3.0.CO;2-4. PMID 9466656.
- ^ Mihaly SR, Ninomiya-Tsuji J, Morioka S (November 2014). "TAK1 control of cell death". Cell Death and Differentiation. 21 (11): 1667–76. doi:10.1038/cdd.2014.123. PMC 4211365. PMID 25146924.
- ^ "Entrez Gene: MAP3K7 mitogen-activated protein kinase kinase kinase 7".
- ^ Scarneo, Scott, Xin Zhang, Yaomin Wang, Jose Camacho-Domenech, Jennifer Ricano, Philip Hughes, Tim Haystead, and Andrea G. Nackley. "Transforming growth factor-β-activated kinase 1 (TAK1) mediates chronic pain and cytokine production in mouse models of inflammatory, neuropathic, and primary pain." The Journal of Pain (2023).
- ^ Scarneo, S. A., Yang, K. W., Roques, J. R., Dai, A., Eibschutz, L. S., Hughes, P., & Haystead, T. A. (2020). TAK1 regulates the tumor microenvironment through inflammatory, angiogenetic and apoptotic signaling cascades. Oncotarget, 11(21), 1961.
- ^ Scarneo, S. A., Eibschutz, L. S., Bendele, P. J., Yang, K. W., Totzke, J., Hughes, P., ... & Haystead, T. A. (2019). Pharmacological inhibition of TAK1, with the selective inhibitor takinib, alleviates clinical manifestation of arthritis in CIA mice. Arthritis Research & Therapy, 21(1), 1-10.
- ^ Scarneo, S. A., Mansourati, A., Eibschutz, L. S., Totzke, J., Roques, J. R., Loiselle, D., ... & Haystead, T. A. (2018). Genetic and pharmacological validation of TAK1 inhibition in macrophages as a therapeutic strategy to effectively inhibit TNF secretion. Scientific reports, 8(1), 1-11.
- ^ a b Mochida Y, Takeda K, Saitoh M, Nishitoh H, Amagasa T, Ninomiya-Tsuji J, Matsumoto K, Ichijo H (October 2000). "ASK1 inhibits interleukin-1-induced NF-kappa B activity through disruption of TRAF6-TAK1 interaction". J. Biol. Chem. 275 (42): 32747–32752. doi:10.1074/jbc.M003042200. PMID 10921914.
- ^ a b c d Ninomiya-Tsuji J, Kishimoto K, Hiyama A, Inoue J, Cao Z, Matsumoto K (March 1999). "The kinase TAK1 can activate the NIK-I kappaB as well as the MAP kinase cascade in the IL-1 signalling pathway". Nature. 398 (6724): 252–256. Bibcode:1999Natur.398..252N. doi:10.1038/18465. PMID 10094049. S2CID 4421236.
- ^ Sakurai H, Miyoshi H, Toriumi W, Sugita T (April 1999). "Functional interactions of transforming growth factor beta-activated kinase 1 with IkappaB kinases to stimulate NF-kappaB activation". J. Biol. Chem. 274 (15): 10641–10648. doi:10.1074/jbc.274.15.10641. PMID 10187861.
- ^ a b Sakurai H, Miyoshi H, Mizukami J, Sugita T (June 2000). "Phosphorylation-dependent activation of TAK1 mitogen-activated protein kinase kinase kinase by TAB1". FEBS Lett. 474 (2–3): 141–145. doi:10.1016/s0014-5793(00)01588-x. PMID 10838074. S2CID 30831863.
- ^ a b c Ishitani T, Takaesu G, Ninomiya-Tsuji J, Shibuya H, Gaynor RB, Matsumoto K (December 2003). "Role of the TAB2-related protein TAB3 in IL-1 and TNF signaling". EMBO J. 22 (23): 6277–6288. doi:10.1093/emboj/cdg605. PMC 291846. PMID 14633987.
- ^ Wang C, Deng L, Hong M, Akkaraju GR, Inoue J, Chen ZJ (July 2001). "TAK1 is a ubiquitin-dependent kinase of MKK and IKK". Nature. 412 (6844): 346–351. Bibcode:2001Natur.412..346W. doi:10.1038/35085597. PMID 11460167. S2CID 9641806.
- ^ a b c Cheung PC, Nebreda AR, Cohen P (February 2004). "TAB3, a new binding partner of the protein kinase TAK1". Biochem. J. 378 (Pt 1): 27–34. doi:10.1042/BJ20031794. PMC 1223947. PMID 14670075.
- ^ Channavajhala PL, Wu L, Cuozzo JW, Hall JP, Liu W, Lin LL, Zhang Y (November 2003). "Identification of a novel human kinase supporter of Ras (hKSR-2) that functions as a negative regulator of Cot (Tpl2) signaling". J. Biol. Chem. 278 (47): 47089–47097. doi:10.1074/jbc.M306002200. PMID 12975377.
- ^ a b Takaesu G, Kishida S, Hiyama A, Yamaguchi K, Shibuya H, Irie K, Ninomiya-Tsuji J, Matsumoto K (April 2000). "TAB2, a novel adaptor protein, mediates activation of TAK1 MAPKKK by linking TAK1 to TRAF6 in the IL-1 signal transduction pathway". Mol. Cell. 5 (4): 649–658. doi:10.1016/s1097-2765(00)80244-0. PMID 10882101.
- ^ a b Takaesu G, Ninomiya-Tsuji J, Kishida S, Li X, Stark GR, Matsumoto K (April 2001). "Interleukin-1 (IL-1) receptor-associated kinase leads to activation of TAK1 by inducing TAB2 translocation in the IL-1 signaling pathway". Mol. Cell. Biol. 21 (7): 2475–2484. doi:10.1128/MCB.21.7.2475-2484.2001. PMC 86880. PMID 11259596.
- ^ Kimura N, Matsuo R, Shibuya H, Nakashima K, Taga T (June 2000). "BMP2-induced apoptosis is mediated by activation of the TAK1-p38 kinase pathway that is negatively regulated by Smad6". J. Biol. Chem. 275 (23): 17647–17652. doi:10.1074/jbc.M908622199. PMID 10748100.
- ^ Yanagisawa M, Nakashima K, Takeda K, Ochiai W, Takizawa T, Ueno M, Takizawa M, Shibuya H, Taga T (December 2001). "Inhibition of BMP2-induced, TAK1 kinase-mediated neurite outgrowth by Smad6 and Smad7". Genes Cells. 6 (12): 1091–1099. doi:10.1046/j.1365-2443.2001.00483.x. PMID 11737269. S2CID 25476125.
- ^ Hanada M, Ninomiya-Tsuji J, Komaki K, Ohnishi M, Katsura K, Kanamaru R, Matsumoto K, Tamura S (February 2001). "Regulation of the TAK1 signaling pathway by protein phosphatase 2C". J. Biol. Chem. 276 (8): 5753–5759. doi:10.1074/jbc.M007773200. PMID 11104763.
- ^ Walsh MC, Kim GK, Maurizio PL, Molnar EE, Choi Y (2008). Unutmaz D (ed.). "TRAF6 autoubiquitination-independent activation of the NFkappaB and MAPK pathways in response to IL-1 and RANKL". PLOS ONE. 3 (12): e4064. Bibcode:2008PLoSO...3.4064W. doi:10.1371/journal.pone.0004064. PMC 2603309. PMID 19112497.
- ^ Sorrentino A, Thakur N, Grimsby S, Marcusson A, von Bulow V, Schuster N, Zhang S, Heldin CH, Landström M (October 2008). "The type I TGF-beta receptor engages TRAF6 to activate TAK1 in a receptor kinase-independent manner". Nat. Cell Biol. 10 (10): 1199–1207. doi:10.1038/ncb1780. PMID 18758450. S2CID 22984417.
Further reading
[edit]- Acton A (2012). Transforming Growth Factors: Advances in Research and Application: 2011 Edition. Vol. 1. ScholarlyEditions. pp. 1–151. ISBN 9781464927331.
- Lin A (2006). The JNK Signaling Pathway (Molecular Biology Intelligence Unit). Vol. 1. Landes Bioscience. pp. 1–97. ISBN 978-1587061202.
- Karin M (2011). NF-kB in Health and Disease. Current Topics in Microbiology and Immunology. Vol. 1. Springer Science & Business Media. pp. 1–268. doi:10.1007/978-3-642-16017-2. ISBN 978-3642160165.
- Wu H (2007). TNF Receptor Associated Factors (TRAFs). Vol. 1. Springer Science & Business Media. pp. 1–206. ISBN 978-0397507597.
External links
[edit]- Overview of all the structural information available in the PDB for UniProt: O43318 (Mitogen-activated protein kinase kinase kinase 7) at the PDBe-KB.
| |||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||
| Activity | |
|---|---|
| Regulation | |
| Classification | |
| Kinetics | |
| Types |
|